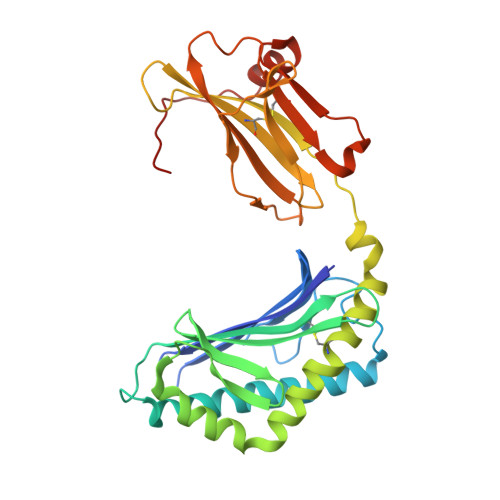
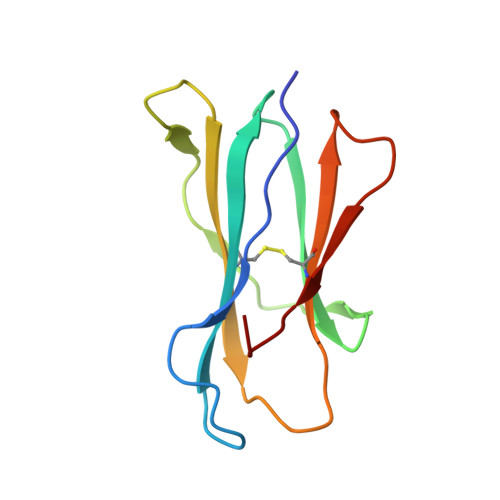
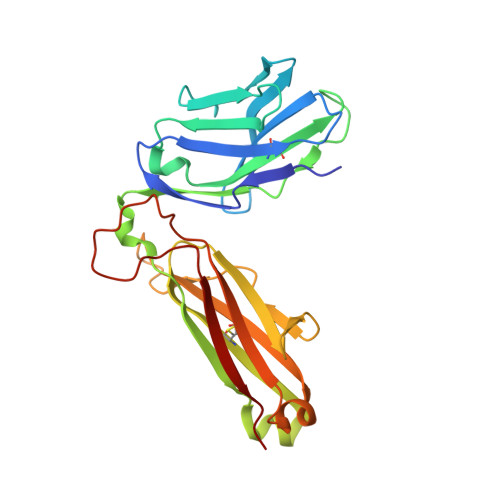
Recognition of CD1d-sulfatide mediated by a type II natural killer T cell antigen receptor.
Patel, O., Pellicci, D.G., Gras, S., Sandoval-Romero, M.L., Uldrich, A.P., Mallevaey, T., Clarke, A.J., Le Nours, J., Theodossis, A., Cardell, S.L., Gapin, L., Godfrey, D.I., Rossjohn, J.(2012) Nat Immunol 13: 857-863
- PubMed: 22820603 Search on PubMed
- DOI: https://doi.org/10.1038/ni.2372
- Primary Citation Related Structures:
4EI5, 4EI6 - PubMed Abstract:
Natural killer T cells (NKT cells) are divided into type I and type II subsets on the basis of differences in their T cell antigen receptor (TCR) repertoire and CD1d-antigen specificity. Although the mode by which type I NKT cell TCRs recognize CD1d-antigen has been established, how type II NKT cell TCRs engage CD1d-antigen is unknown. Here we provide a basis for how a type II NKT cell TCR, XV19, recognized CD1d-sulfatide. The XV19 TCR bound orthogonally above the A' pocket of CD1d, in contrast to the parallel docking of type I NKT cell TCRs over the F' pocket of CD1d. At the XV19 TCR-CD1d-sulfatide interface, the TCRα and TCRβ chains sat centrally on CD1d, where the malleable CDR3 loops dominated interactions with CD1d-sulfatide. Accordingly, we highlight the diverse mechanisms by which NKT cell TCRs can bind CD1d and account for the distinct antigen specificity of type II NKT cells.
- 1] Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Australia.
Organizational Affiliation: