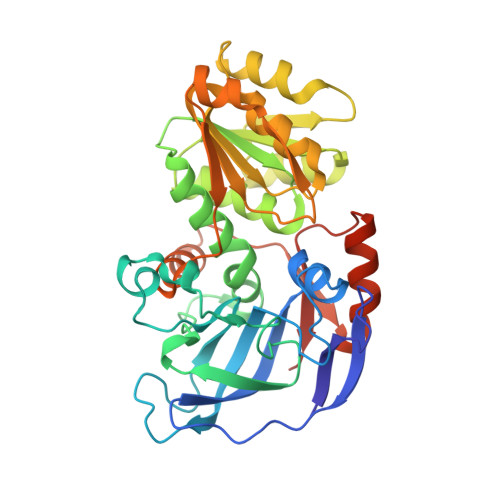
Structure-guided engineering of Lactococcus lactis alcohol dehydrogenase LlAdhA for improved conversion of isobutyraldehyde to isobutanol.
Liu, X., Bastian, S., Snow, C.D., Brustad, E.M., Saleski, T.E., Xu, J.H., Meinhold, P., Arnold, F.H.(2012) J Biotechnol 164: 188-195
- PubMed: 22974724 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbiotec.2012.08.008
- Primary Citation Related Structures:
4EEX, 4EEZ - PubMed Abstract:
We have determined the X-ray crystal structures of the NADH-dependent alcohol dehydrogenase LlAdhA from Lactococcus lactis and its laboratory-evolved variant LlAdhA(RE1) at 1.9Å and 2.5Å resolution, respectively. LlAdhA(RE1), which contains three amino acid mutations (Y50F, I212T, and L264V), was engineered to increase the microbial production of isobutanol (2-methylpropan-1-ol) from isobutyraldehyde (2-methylpropanal). Structural comparison of LlAdhA and LlAdhA(RE1) indicates that the enhanced activity on isobutyraldehyde stems from increases in the protein's active site size, hydrophobicity, and substrate access. Further structure-guided mutagenesis generated a quadruple mutant (Y50F/N110S/I212T/L264V), whose KM for isobutyraldehyde is ∼17-fold lower and catalytic efficiency (kcat/KM) is ∼160-fold higher than wild-type LlAdhA. Combining detailed structural information and directed evolution, we have achieved significant improvements in non-native alcohol dehydrogenase activity that will facilitate the production of next-generation fuels such as isobutanol from renewable resources.
- Division of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena, CA 91125, USA.
Organizational Affiliation: