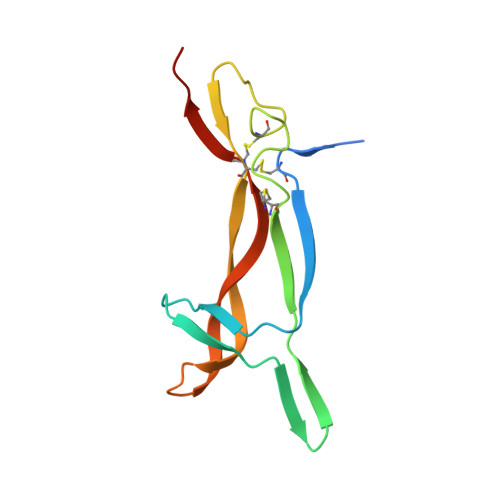
Structural and functional insights into lipid-bound nerve growth factors
Tong, Q., Wang, F., Zhou, H.Z., Sun, H.L., Song, H., Shu, Y.Y., Gong, Y., Zhang, W.T., Cai, T.X., Yang, F.Q., Tang, J., Jiang, T.(2012) FASEB J 26: 3811-3821
- PubMed: 22649032 Search on PubMed
- DOI: https://doi.org/10.1096/fj.12-207316
- Primary Citation Related Structures:
4EAX, 4EC7 - PubMed Abstract:
Nerve growth factor (NGF) is a dimeric molecule that modulates the survival, proliferation, and differentiation of nervous cells and is also known to act on cells of the immune system and endocrine system. NGFs extracted from mouse submaxillary gland and cobra venom have different immunological behaviors, yet the underlying mechanism remains unclear. Here we report the crystal structure of the NGF purified from Chinese cobra Naja naja atra (cNGF), which unexpectedly reveals a 2-tailed lipid molecule that is embedded between the two protomers of the NGF homodimer. In addition, crystallographic analysis indicated that the purified mouse NGF(mNGF) is free from lipid but can bind lysophosphatidylserine (lyso-PS) in the same pocket as cNGF. Bioassays indicated that the binding of lipid molecules to cNGF and mNGF are essential for their mast cell activation activity and abates their p75(NTR) binding capacity. Taken together, these results suggest a new mechanism for the regulation of the function of NGF.
- National Key Laboratory of Biomacromolecules, Institute of Biophysics, Beijing, China.
Organizational Affiliation: