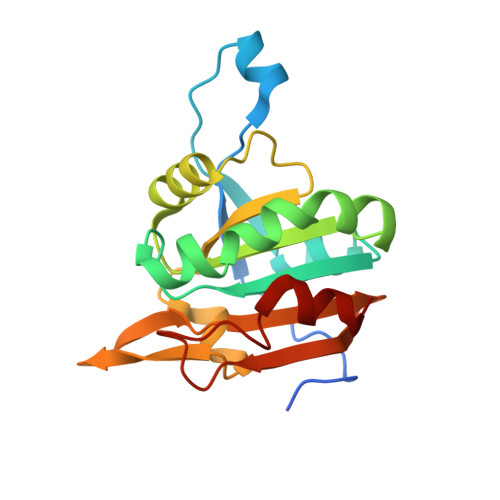
Structural Snapshots of Yeast Alkyl Hydroperoxide Reductase Ahp1 Peroxiredoxin Reveal a Novel Two-cysteine Mechanism of Electron Transfer to Eliminate Reactive Oxygen Species.
Lian, F.M., Yu, J., Ma, X.X., Yu, X.J., Chen, Y., Zhou, C.Z.(2012) J Biological Chem 287: 17077-17087
- PubMed: 22474296 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.357368
- Primary Citation Related Structures:
4DSQ, 4DSR, 4DSS - PubMed Abstract:
Peroxiredoxins (Prxs) are thiol-specific antioxidant proteins that protect cells against reactive oxygen species and are involved in cellular signaling pathways. Alkyl hydroperoxide reductase Ahp1 belongs to the Prx5 subfamily and is a two-cysteine (2-Cys) Prx that forms an intermolecular disulfide bond. Enzymatic assays and bioinformatics enabled us to re-assign the peroxidatic cysteine (C(P)) to Cys-62 and the resolving cysteine (C(R)) to Cys-31 but not the previously reported Cys-120. Thus Ahp1 represents the first 2-Cys Prx with a peroxidatic cysteine after the resolving cysteine in the primary sequence. We also found the positive cooperativity of the substrate t-butyl hydroperoxide binding to Ahp1 homodimer at a Hill coefficient of ∼2, which enabled Ahp1 to eliminate hydroperoxide at much higher efficiency. To gain the structural insights into the catalytic cycle of Ahp1, we determined the crystal structures of Ahp1 in the oxidized, reduced, and Trx2-complexed forms at 2.40, 2.91, and 2.10 Å resolution, respectively. Structural superposition of the oxidized to the reduced form revealed significant conformational changes at the segments containing C(P) and C(R). An intermolecular C(P)-C(R) disulfide bond crossing the A-type dimer interface distinguishes Ahp1 from other typical 2-Cys Prxs. The structure of the Ahp1-Trx2 complex showed for the first time how the electron transfers from thioredoxin to a peroxidase with a thioredoxin-like fold. In addition, site-directed mutagenesis in combination with enzymatic assays suggested that the peroxidase activity of Ahp1 would be altered upon the urmylation (covalently conjugated to ubiquitin-related modifier Urm1) of Lys-32.
- Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, 230027, China.
Organizational Affiliation: