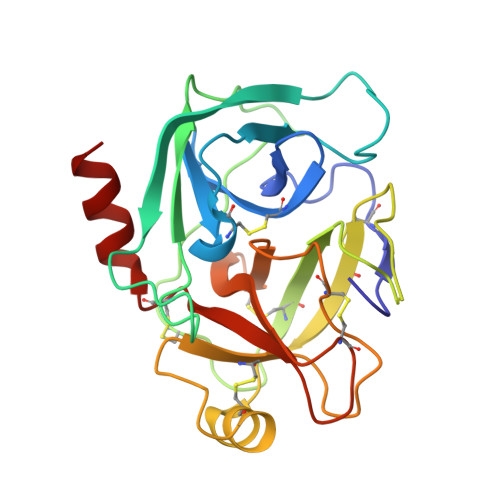
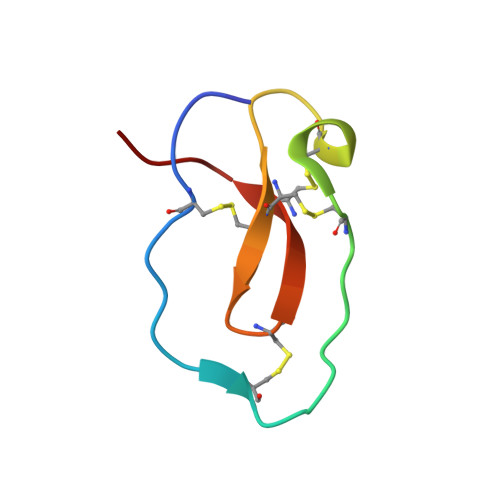
Structure basis 1/2SLPI and porcine pancreas trypsin interaction
Fukushima, K., Kamimura, T., Takimoto-Kamimura, M.(2013) J Synchrotron Radiat 20: 943-947
- PubMed: 24121345 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S090904951302133X
- Primary Citation Related Structures:
4DOQ - PubMed Abstract:
SLPI (secretory leukocyte protease inhibitor) is a 107-residue protease inhibitor which inhibits various serine proteases, including elastase, cathepsin G, chymotrypsin and trypsin. SLPI is obtained as a multiple inhibitor in lung defense and in chronic airway infection. X-ray crystal structures have so far reported that they are full-length SLPIs with bovine α-chymotrypsin and 1/2SLPI (recombinant C-terminal domain of SLPI; Arg58-Ala107) with HNE (human neutrophil elastase). To understand the role of this multiple inhibitory mechanism, the crystal structure of 1/2SLPI with porcine pancreas trypsin was solved and the binding modes of two other complexes compared. The Leu residue surprisingly interacts with the S1 site of trypsin, as with chymotrypsin and elastase. The inhibitory mechanism of 1/2SLPI using the wide primary binding site contacts (from P2' to P5) with various serine proteases is discussed. These inhibitory mechanisms have been acquired in the evolution of the protection system for acute inflammatory diseases.
- Medicinal Chemistry Technology Department, Teijin Institute for Bio-Medical Research, 4-3-2 Asahigaoka, Hino-shi, Tokyo 191-8512, Japan.
Organizational Affiliation: