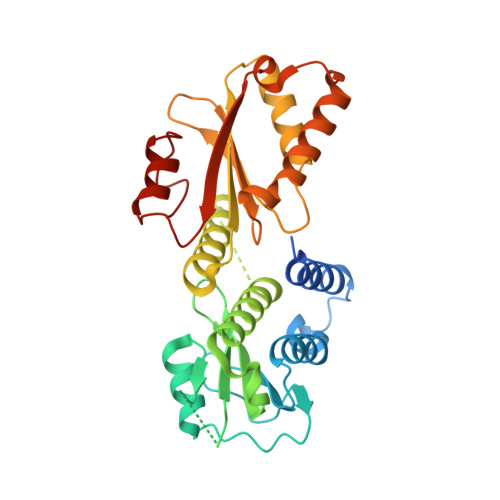
Structure of the Cytoplasmic Region of PelD, a Degenerate Diguanylate Cyclase Receptor That Regulates Exopolysaccharide Production in Pseudomonas aeruginosa.
Whitney, J.C., Colvin, K.M., Marmont, L.S., Robinson, H., Parsek, M.R., Howell, P.L.(2012) J Biological Chem 287: 23582-23593
- PubMed: 22605337 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.375378
- Primary Citation Related Structures:
4DMZ, 4DN0 - PubMed Abstract:
High cellular concentrations of bis-(3',5')-cyclic dimeric guanosine mono-phosphate (c-di-GMP) regulate a diverse range of phenotypes in bacteria including biofilm development. The opportunistic pathogen Pseudomonas aeruginosa produces the PEL polysaccharide to form a biofilm at the air-liquid interface of standing cultures. Among the proteins required for PEL polysaccharide production, PelD has been identified as a membrane-bound c-di-GMP-specific receptor. In this work, we present the x-ray crystal structure of a soluble cytoplasmic region of PelD in its apo and c-di-GMP complexed forms. The structure of PelD reveals an N-terminal GAF domain and a C-terminal degenerate GGDEF domain, the latter of which binds dimeric c-di-GMP at an RXXD motif that normally serves as an allosteric inhibition site for active diguanylate cyclases. Using isothermal titration calorimetry, we demonstrate that PelD binds c-di-GMP with low micromolar affinity and that mutation of residues involved in binding not only decreases the affinity of this interaction but also abrogates PEL-specific phenotypes in vivo. Bioinformatics analysis of the juxtamembrane region of PelD suggests that it contains an α-helical stalk region that connects the soluble region to the transmembrane domains and that similarly to other GAF domain containing proteins, this region likely forms a coiled-coil motif that mediates dimerization. PelD with Alg44 and BcsA of the alginate and cellulose secretion systems, respectively, collectively constitute a group of c-di-GMP receptors that appear to regulate exopolysaccharide assembly at the protein level through activation of their associated glycosyl transferases.
- Program in Molecular Structure and Function, Hospital for Sick Children, Toronto, Ontario M5G 1X8, Canada.
Organizational Affiliation: