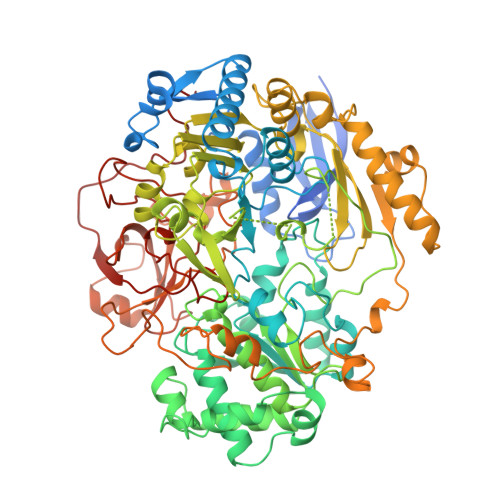
The high resolution crystal structure of DMSO reductase in complex with DMSO.
McAlpine, A.S., McEwan, A.G., Bailey, S.(1998) J Mol Biology 275: 613-623
- PubMed: 9466935 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1513
- Primary Citation Related Structures:
4DMR - PubMed Abstract:
The crystal structure of the molybdenum enzyme dimethylsulphoxide reductase (DMSOR) has been determined at 1.9 A resolution with substrate bound at the active site. DMSOR is an oxotransferase which catalyses the reduction of dimethylsulphoxide (DMSO) to dimethylsulphide (DMS) in a two stage reaction which is linked to oxygen atom transfer and electron transfer. In the first step, DMSO binds to reduced (Mo(IV)) enzyme, the enzyme is oxidised to Mo(VI) with an extra oxygen ligand and DMS is released. Regeneration of reduced enzyme is achieved by transfer of two electrons, successively from a specific cytochrome, and release of the oxygen as water. The enzyme, purified under aerobic conditions, is in the oxidised (Mo(VI)) state. Addition of a large excess of DMS to the oxidised enzyme in solution causes a change in the absorption spectrum of the enzyme. The same reaction occurs within crystals of the enzyme and the crystal structure reveals a complex with DMSO bound to the molybdenum via its oxygen atom. X-ray edge data indicate that the metal is in the Mo(IV) state. The DMSO ligand replaces one of the two oxo groups which ligate the oxidised form of the enzyme, suggesting very strongly that this is the oxygen which is transferred during catalysis. Residues 384 to 390, disordered in the oxidised enzyme, are now clearly seen in the cleft leading to the active site. The side-chain of Trp388 forms a lid trapping the substrate/product.
- CCLRC Daresbury Laboratory, Cheshire, UK.
Organizational Affiliation: