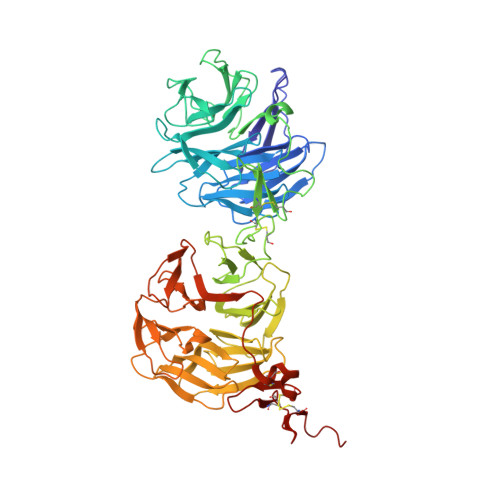
Characterization of the interaction of sclerostin with the LRP family of Wnt co-receptors
Holdsworth, G., Slocombe, P., Doyle, C., Sweeney, B., Veverka, V., Le Rich, K., Franklin, R.J., Brookings, D., Turner, J., Kennedy, J., Garlish, R., Shi, J., Newnham, L., McMillan, D., Muzylak, M., Carr, M., Henry, A.J., Ceska, T., Robinson, M.K.To be published.