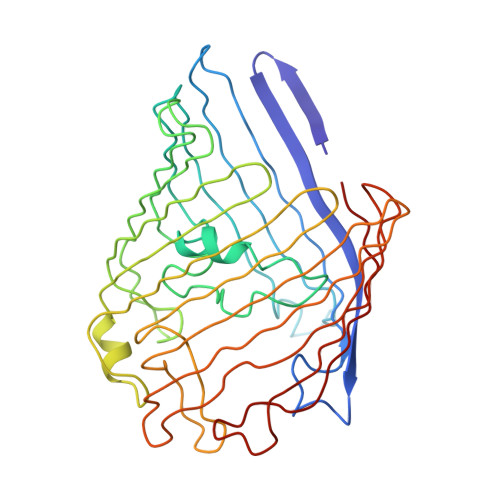
Two Different Centered Monoclinic Crystals of the E. Coli Outer-Membrane Protein Ompf Originate from the Same Building Block.
Chaptal, V., Kilburg, A., Flot, D., Wiseman, B., Aghajari, N., Jault, J., Falson, P.(2016) Biochim Biophys Acta 1858: 326
- PubMed: 26620074 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbamem.2015.11.021
- Primary Citation Related Structures:
4D5U - PubMed Abstract:
Macromolecule crystal formation can be divided in two major steps: 1. the formation of a nucleus and 2. the growth of this nucleus into a full mature crystal. The latter is well described and understood, while the former remains elusive due to the difficulty to study it and is described by nucleation theories. Here we report the structure of the Escherichia coli outer membrane porin OmpF in two centered monoclinic space groups. Strikingly, the two crystals originate from the same building block, made of two trimers of OmpF interacting via their rough side. The different crystallization conditions trigger the formation of distinct arrangement of these building blocks, leading to the formation of translational non-crystallographic symmetry (tNCS) in one case, made possible by the loose lateral packing mediated by detergents. In light of nucleation theories, these results allow us to speculate that these two crystals originate from nuclei made of either clusters of building blocks, or already forming columns that later associate laterally using detergents as glue.
- Drug Resistance Mechanism and Modulation team, Institut de Biologie et Chimie des Protéines (IBCP), UMR5086 CNRS/Université Lyon 1, 7 passage du Vercors, F-69367 Lyon, France. Electronic address: vincent.chaptal@ibcp.fr.
Organizational Affiliation: