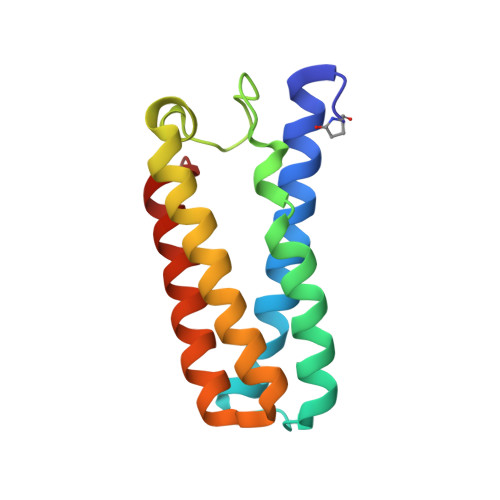
Hydrogen Bonding of the Dissociated Histidine Ligand is not Required for Formation of a Proximal No Adduct in Cytochrome C'.
Ghafoor, D.D., Kekilli, D., Abdullah, G.H., Dworkowski, F.S.N., Hassan, H.G., Wilson, M.T., Strange, R.W., Hough, M.A.(2015) J Biol Inorg Chem 20: 949
- PubMed: 26100643 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-015-1278-y
- Primary Citation Related Structures:
4D4N, 4D4X, 5AGF - PubMed Abstract:
Cytochromes c', that occur in methanotrophic, denitrifying and photosynthetic bacteria, form unusual proximal penta-coordinate NO complexes via a hexa-coordinate distal NO intermediate. Their NO binding properties are similar to those of the eukaryotic NO sensor, soluble guanylate cyclase, for which they provide a valuable structural model. Previous studies suggested that hydrogen bonding between the displaced proximal histidine (His120) ligand (following its dissociation from heme due to trans effects from the distally bound NO) and a conserved aspartate residue (Asp121) could play a key role in allowing proximal NO binding to occur. We have characterized three variants of Alcaligenes xylosoxidans cytochrome c' (AXCP) where Asp121 has been replaced by Ala, Ile and Gln, respectively. In all variants, hydrogen bonding between residue 121 and His120 is abolished yet 5-coordinate proximal NO species are still formed. Our data therefore demonstrate that the His120-Asp121 bond is not essential for proximal NO binding although it likely provides an energy minimum for the displaced His ligand. All variants have altered proximal pocket structure relative to native AXCP.
- Faculty of Science and Education Science, University of Sulaimani, Sulaymaniyah, Iraq.
Organizational Affiliation: