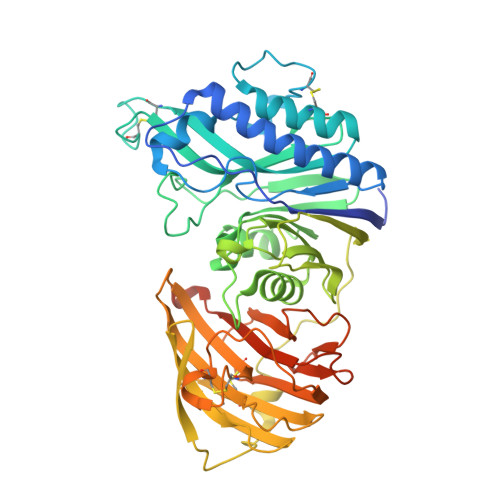
The Structure of Vanin-1: A Key Enzyme Linking Metabolic Disease and Inflammation
Boersma, Y.L., Newman, J., Adams, T.E., Cowieson, N., Krippner, G., Bozaoglu, K., Peat, T.S.(2014) Acta Crystallogr D Biol Crystallogr 70: 3320
- PubMed: 25478849 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004714022767
- Primary Citation Related Structures:
4CYF, 4CYG, 4CYY - PubMed Abstract:
Although part of the coenzyme A pathway, vanin 1 (also known as pantetheinase) sits on the cell surface of many cell types as an ectoenzyme, catalyzing the breakdown of pantetheine to pantothenic acid (vitamin B5) and cysteamine, a strong reducing agent. Vanin 1 was initially discovered as a protein involved in the homing of leukocytes to the thymus. Numerous studies have shown that vanin 1 is involved in inflammation, and more recent studies have shown a key role in metabolic disease. Here, the X-ray crystal structure of human vanin 1 at 2.25 Å resolution is presented, which is the first reported structure from the vanin family, as well as a crystal structure of vanin 1 bound to a specific inhibitor. These structures illuminate how vanin 1 can mediate its biological roles by way of both enzymatic activity and protein-protein interactions. Furthermore, it sheds light on how the enzymatic activity is regulated by a novel allosteric mechanism at a domain interface.
- Department of Pharmaceutical Biology, Groningen Research Institute of Pharmacy, University of Groningen, A. Deusinglaan 1, 9713 AV Groningen, The Netherlands.
Organizational Affiliation: