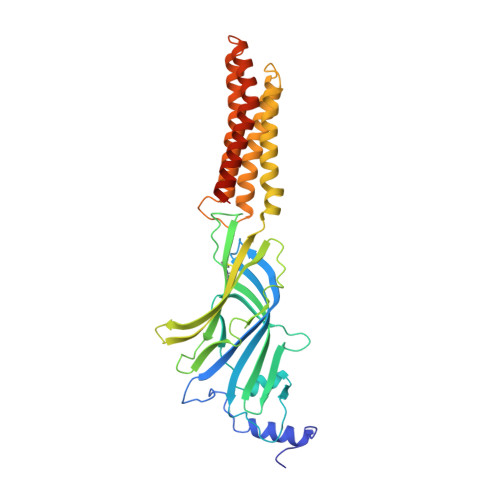
Crystal Structure of a Human Gabaa Receptor
Miller, P.S., Aricescu, A.R.(2014) Nature 512: 270
- PubMed: 24909990 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature13293
- Primary Citation Related Structures:
4COF - PubMed Abstract:
Type-A γ-aminobutyric acid receptors (GABAARs) are the principal mediators of rapid inhibitory synaptic transmission in the human brain. A decline in GABAAR signalling triggers hyperactive neurological disorders such as insomnia, anxiety and epilepsy. Here we present the first three-dimensional structure of a GABAAR, the human β3 homopentamer, at 3 Å resolution. This structure reveals architectural elements unique to eukaryotic Cys-loop receptors, explains the mechanistic consequences of multiple human disease mutations and shows an unexpected structural role for a conserved N-linked glycan. The receptor was crystallized bound to a previously unknown agonist, benzamidine, opening a new avenue for the rational design of GABAAR modulators. The channel region forms a closed gate at the base of the pore, representative of a desensitized state. These results offer new insights into the signalling mechanisms of pentameric ligand-gated ion channels and enhance current understanding of GABAergic neurotransmission.
- Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Roosevelt Drive, Oxford OX3 7BN, UK.
Organizational Affiliation: