Plasmepsin Inhibitory Activity and Structure-Guided Optimization of a Potent Hydroxyethylamine-Based Antimalarial Hit.
Jaudzems, K., Tars, K., Maurops, G., Ivdra, N., Otikovs, M., Leitans, J., Kanepe-Lapsa, I., Domraceva, I., Mutule, I., Trapencieris, P., Blackman, M.J., Jirgensons, A.(2014) ACS Med Chem Lett 5: 373
- PubMed: 24900843 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml4004952
- Primary Citation Related Structures:
4CKU - PubMed Abstract:
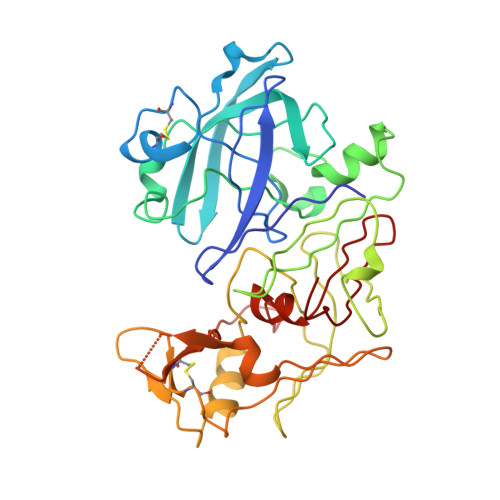
Antimalarial hit 1 SR (TCMDC-134674) identified in a GlaxoSmithKline cell based screening campaign was evaluated for inhibitory activity against the digestive vacuole plasmepsins (Plm I, II, and IV). It was found to be a potent Plm IV inhibitor with no selectivity over Cathepsin D. A cocrystal structure of 1 SR bound to Plm II was solved, providing structural insight for the design of more potent and selective analogues. Structure-guided optimization led to the identification of structurally simplified analogues 17 and 18 as low nanomolar inhibitors of both, plasmepsin Plm IV activity and P. falciparum growth in erythrocytes.
- Latvian Institute of Organic Synthesis , Aizkraukles 21, Riga LV-1006, Latvia.
Organizational Affiliation: