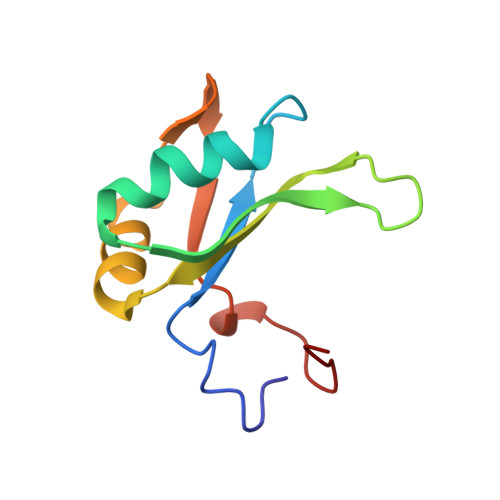
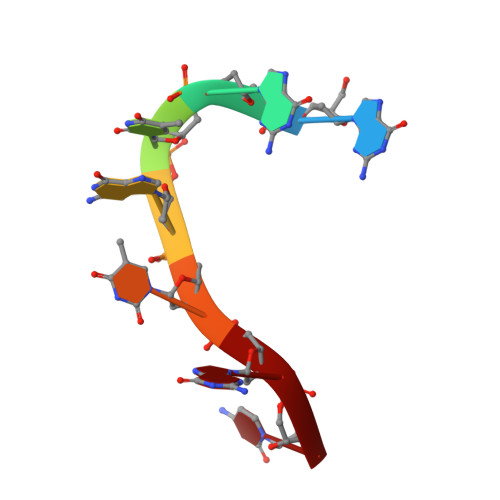
Backbone-Independent Nucleic Acid Binding by Splicing Factor Sup-12 Reveals Key Aspects of Molecular Recognition
Amrane, S., Rebora, K., Zniber, I., Dupuy, D., Mackereth, C.D.(2014) Nat Commun 5: 4595
- PubMed: 25183497 Search on PubMed
- DOI: https://doi.org/10.1038/ncomms5595
- Primary Citation Related Structures:
4CH0, 4CH1, 4CIO - PubMed Abstract:
Cellular differentiation is frequently accompanied by alternative splicing, enabled by the expression of tissue-specific factors which bind to pre-mRNAs and regulate exon choice. During Caenorhabditis elegans development, muscle-specific expression of the splicing factor SUP-12, together with a member of the Fox-1 family of splicing proteins, generates a functionally distinct isoform of the fibroblast growth factor receptor EGL-15. Using a combination of NMR spectroscopy and isothermal titration calorimetry, we determined the mode of nucleic acid binding by the RNA recognition motif domain of SUP-12. The calculated structures provide the first atomic details of RNA and DNA binding by the family of proteins that include SUP-12, RBM24, RBM38/RNPC1, SEB-4 and XSeb4R. This information was further used to design strategic mutations to probe the interaction with ASD-1 and to quantitatively perturb splicing in vivo.
- 1] Institut Européen de Chimie et Biologie, IECB, Univ. Bordeaux, 2 rue Robert Escarpit, F-33607 Pessac, France [2] Inserm, U869, ARNA Laboratory, 146 rue Léo Saignat, F-33076 Bordeaux, France.
Organizational Affiliation: