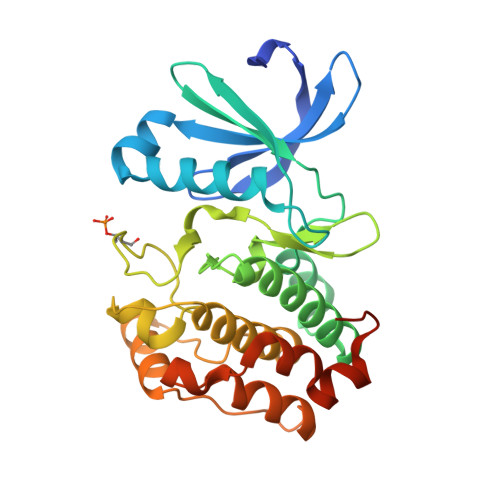
The Structure of C290A:C393A Aurora a Provides Structural Insights Into Kinase Regulation.
Burgess, S.G., Bayliss, R.(2015) Acta Crystallogr Sect F Struct Biol Cryst Commun 71: 315
- PubMed: 25760707 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X15002290
- Primary Citation Related Structures:
4CEG - PubMed Abstract:
Aurora A is a Ser/Thr protein kinase that functions in cell-cycle regulation and is implicated in cancer development. During mitosis, Aurora A is activated by autophosphorylation on its activation loop at Thr288. The Aurora A catalytic domain (amino acids 122-403) expressed in Escherichia coli autophosphorylates on two activation-loop threonine residues (Thr288 and Thr287), whereas a C290A,C393A double point mutant of the Aurora A catalytic domain autophosphorylates only on Thr288. The structure of the complex of this mutant with ADP and magnesium was determined to 2.1 Å resolution using molecular replacement. This is an improvement on the existing 2.75 Å resolution structure of the equivalent wild-type complex. The structure confirms that single phosphorylation of the activation loop on Thr288 is insufficient to stabilize a `fully active' conformation of the activation loop in the absence of binding to TPX2.
- Department of Biochemistry, University of Leicester, Lancaster Road, Leicester LE1 9HN, England.
Organizational Affiliation: