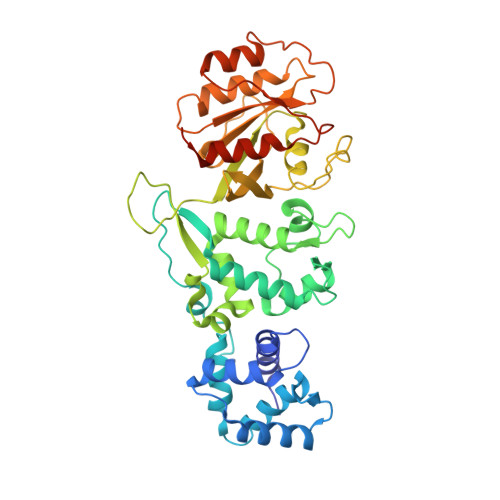
Structural Coupling of the EF Hand and C-Terminal Gtpase Domains in the Mitochondrial Protein Miro.
Klosowiak, J.L., Focia, P.J., Chakravarthy, S., Landahl, E.C., Freymann, D.M., Rice, S.E.(2013) EMBO Rep 14: 968
- PubMed: 24071720 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/embor.2013.151
- Primary Citation Related Structures:
4C0J, 4C0K, 4C0L - PubMed Abstract:
Miro is a highly conserved calcium-binding GTPase at the regulatory nexus of mitochondrial transport and autophagy. Here we present crystal structures comprising the tandem EF hand and carboxy terminal GTPase (cGTPase) domains of Drosophila Miro. The structures reveal two previously unidentified 'hidden' EF hands, each paired with a canonical EF hand. Each EF hand pair is bound to a helix that structurally mimics an EF hand ligand. A key nucleotide-sensing element and a Pink1 phosphorylation site both lie within an extensive EF hand-cGTPase interface. Our results indicate structural mechanisms for calcium, nucleotide and phosphorylation-dependent regulation of mitochondrial function by Miro.
- Department of Cell and Molecular Biology, Feinberg School of Medicine, Northwestern University, 303 East Chicago Avenue, Chicago, Illinois 60611, USA.
Organizational Affiliation: