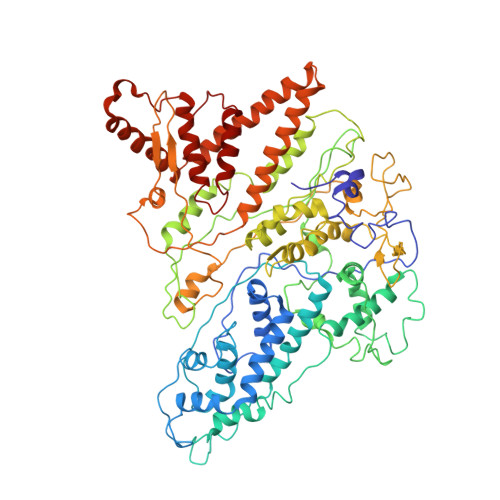
Subunit Folds and Maturation Pathway of a Dsrna Virus Capsid.
Nemecek, D., Boura, E., Wu, W., Cheng, N., Plevka, P., Qiao, J., Mindich, L., Heymann, J.B., Hurley, J.H., Steven, A.C.(2013) Structure 21: 1374
- PubMed: 23891288 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2013.06.007
- Primary Citation Related Structures:
4BTG, 4BTQ, 4K7H - PubMed Abstract:
The cystovirus ϕ6 shares several distinct features with other double-stranded RNA (dsRNA) viruses, including the human pathogen, rotavirus: segmented genomes, nonequivalent packing of 120 subunits in its icosahedral capsid, and capsids as compartments for transcription and replication. ϕ6 assembles as a dodecahedral procapsid that undergoes major conformational changes as it matures into the spherical capsid. We determined the crystal structure of the capsid protein, P1, revealing a flattened trapezoid subunit with an α-helical fold. We also solved the procapsid with cryo-electron microscopy to comparable resolution. Fitting the crystal structure into the procapsid disclosed substantial conformational differences between the two P1 conformers. Maturation via two intermediate states involves remodeling on a similar scale, besides huge rigid-body rotations. The capsid structure and its stepwise maturation that is coupled to sequential packaging of three RNA segments sets the cystoviruses apart from other dsRNA viruses as a dynamic molecular machine.
- National Institute of Arthritis and Musculoskeletal and Skin Diseases, National Institutes of Health, 50 South Drive, Bethesda, MD 20892, USA.
Organizational Affiliation: