Is NMR Fragment Screening Fine-Tuned to Assess Druggability of Protein-Protein Interactions?
Dias, D.M., Van Molle, I., Baud, M.G.J., Galdeano, C., Geraldes, C.F.G.C., Ciulli, A.(2014) ACS Med Chem Lett 5: 23
- PubMed: 24436777 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml400296c
- Primary Citation Related Structures:
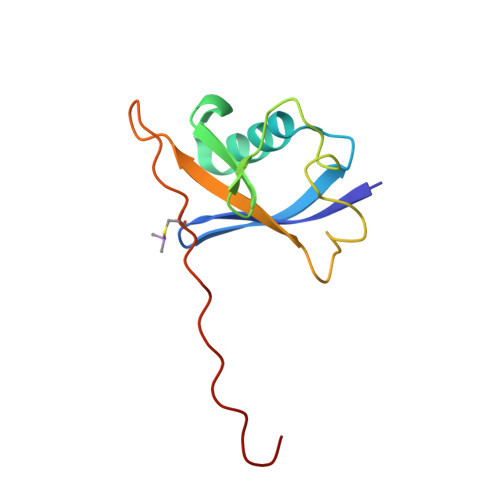
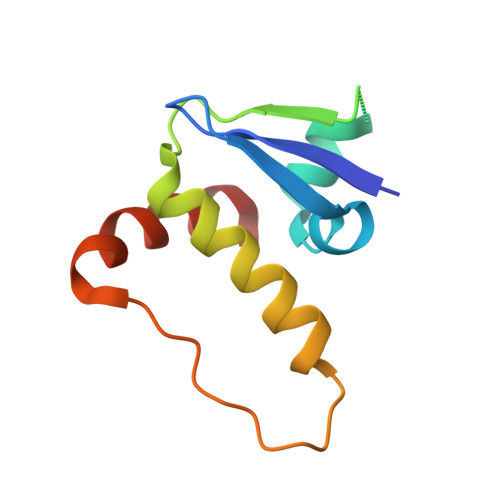
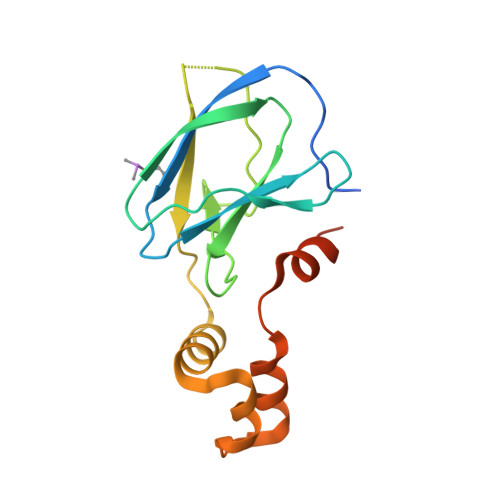
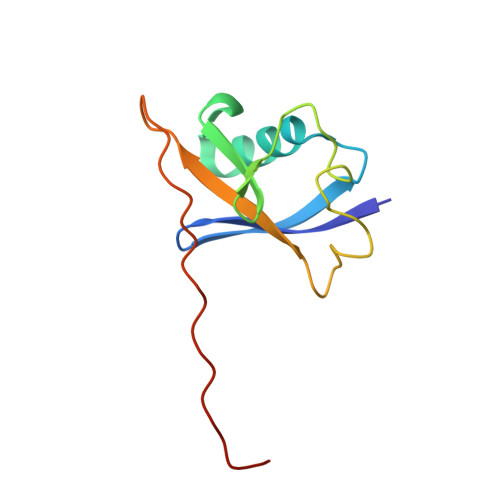
4BKS, 4BKT - PubMed Abstract:
Modulation of protein-protein interactions (PPIs) with small molecules has been hampered by a lack of lucid methods capable of reliably identifying high-quality hits. In fragment screening, the low ligand efficiencies associated with PPI target sites pose significant challenges to fragment binding detection. Here, we investigate the requirements for ligand-based NMR techniques to detect rule-of-three compliant fragments that form part of known high-affinity inhibitors of the PPI between the von Hippel-Lindau protein and the alpha subunit of hypoxia-inducible factor 1 (pVHL:HIF-1α). Careful triaging allowed rescuing weak but specific binding of fragments that would otherwise escape detection at this PPI. Further structural information provided by saturation transfer difference (STD) group epitope mapping, protein-based NMR, competitive isothermal titration calorimetry (ITC), and X-ray crystallography confirmed the binding mode of the rescued fragments. Our findings have important implications for PPI druggability assessment by fragment screening as they reveal an accessible threshold for fragment detection and validation.
- Department of Chemistry, University of Cambridge , Cambridge CB2 1EW, U.K. ; Department of Life Sciences, Faculty of Science and Technology, Centre for Neurosciences and Cell Biology and Chemistry Centre, University of Coimbra , Coimbra, Portugal.
Organizational Affiliation: