ATPase Crystal Structure with Bound Phosphate Analogue
Mattle, D., Drachmann, N.D., Liu, X.Y., Gourdon, P., Pedersen, B.P., Morth, P., Wang, J., Nissen, P.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| COPPER EFFLUX ATPASE | 736 | Legionella pneumophila | Mutation(s): 0 EC: 3.6.3 (PDB Primary Data), 7.2.2.8 (UniProt) Membrane Entity: Yes | 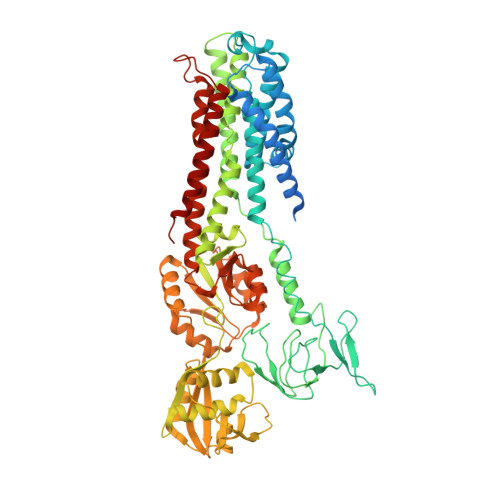 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q5ZWR1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| MGF Download:Ideal Coordinates CCD File | C [auth A] | TRIFLUOROMAGNESATE F3 Mg GJOMWUHGUQLOAC-UHFFFAOYSA-K |  | ||
| MG Download:Ideal Coordinates CCD File | B [auth A] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 44.36 | α = 90 |
| b = 72.65 | β = 90 |
| c = 328.79 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XDS | data scaling |