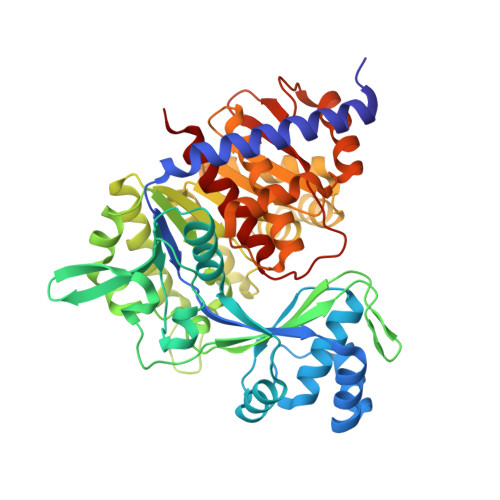
Crystal Structure, Saxs and Kinetic Mechanism of Hyperthermophilic Adp-Dependent Glucokinase from Thermococcus Litoralis Reveal a Conserved Mechanism for Catalysis.
Rivas-Pardo, J.A., Herrera-Morande, A., Castro-Fernandez, V., Fernandez, F.J., Vega, M.C., Guixe, V.(2013) PLoS One 8: 66687
- PubMed: 23818958 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0066687
- Primary Citation Related Structures:
4B8R, 4B8S - PubMed Abstract:
ADP-dependent glucokinases represent a unique family of kinases that belong to the ribokinase superfamily, being present mainly in hyperthermophilic archaea. For these enzymes there is no agreement about the magnitude of the structural transitions associated with ligand binding and whether they are meaningful to the function of the enzyme. We used the ADP-dependent glucokinase from Thermococcus litoralis as a model to investigate the conformational changes observed in X-ray crystallographic structures upon substrate binding and to compare them with those determined in solution in order to understand their interplay with the glucokinase function. Initial velocity studies indicate that catalysis follows a sequential ordered mechanism that correlates with the structural transitions experienced by the enzyme in solution and in the crystal state. The combined data allowed us to resolve the open-closed conformational transition that accounts for the complete reaction cycle and to identify the corresponding clusters of aminoacids residues responsible for it. These results provide molecular bases for a general mechanism conserved across the ADP-dependent kinase family.
- Departamento de Biología, Facultad de Ciencias, Universidad de Chile, Santiago, Chile.
Organizational Affiliation: