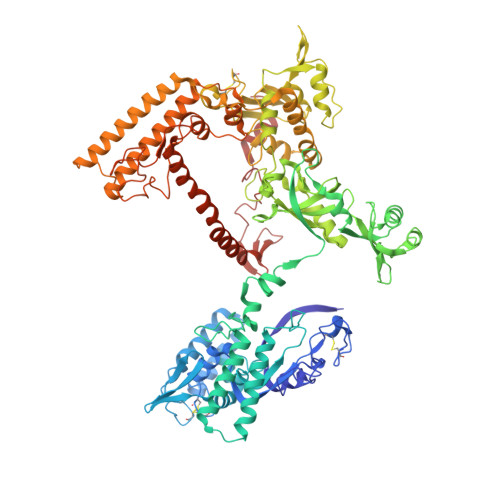
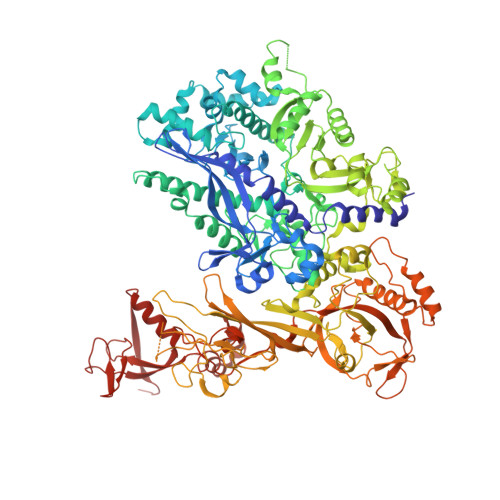
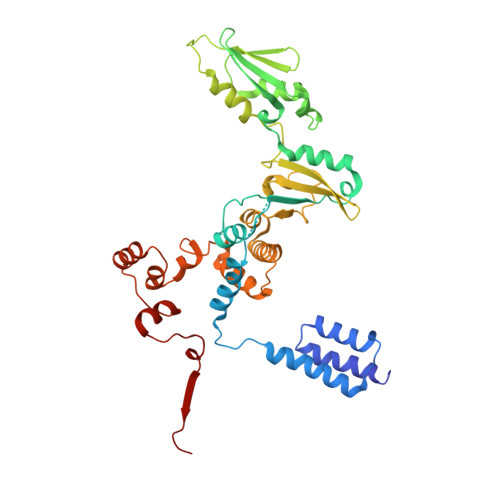
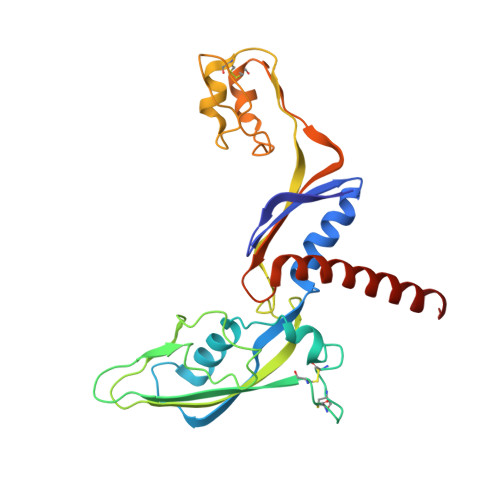
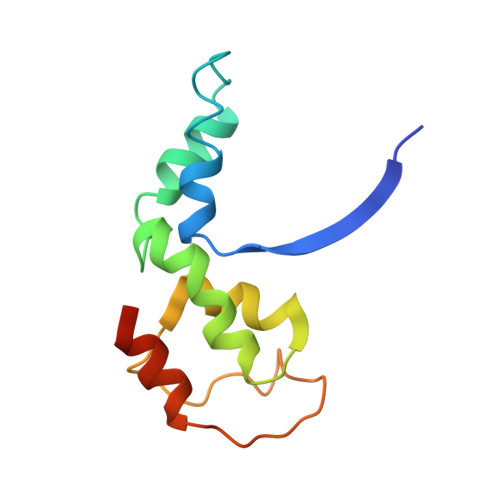
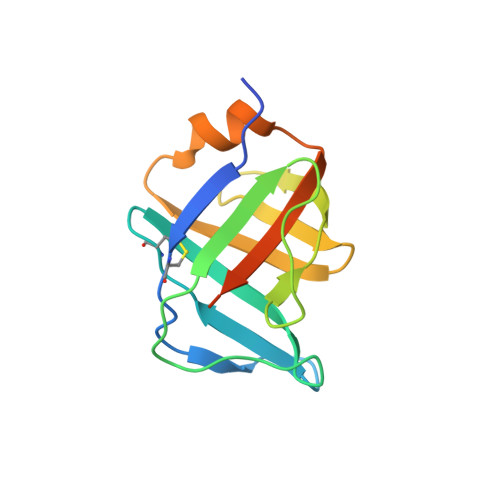
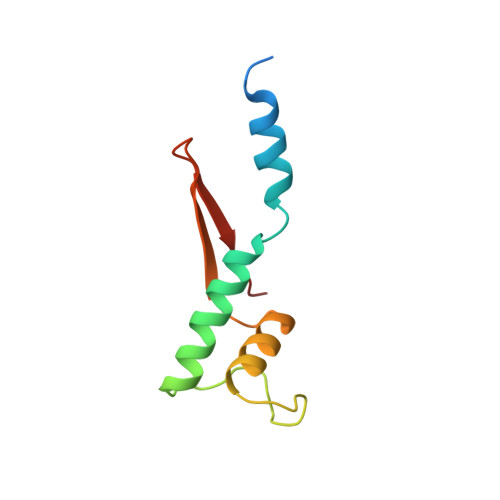
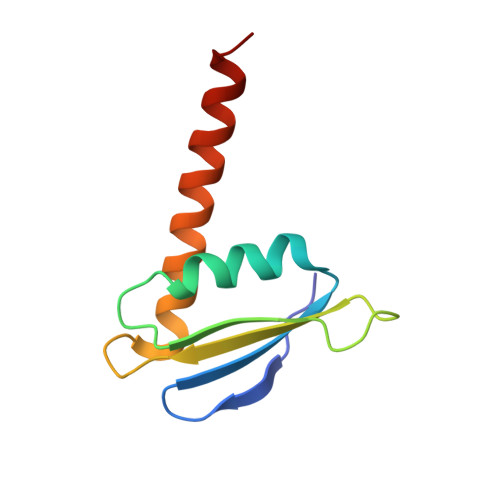
Structural and Functional Analyses of the Interaction of Archaeal RNA Polymerase with DNA.
Wojtas, M.N., Mogni, M., Millet, O., Bell, S.D., Abrescia, N.G.A.(2012) Nucleic Acids Res 40: 9941
- PubMed: 22848102 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gks692
- Primary Citation Related Structures:
4AYB, 4V8S - PubMed Abstract:
Multi-subunit RNA polymerases (RNAPs) in all three domains of life share a common ancestry. The composition of the archaeal RNAP (aRNAP) is not identical between phyla and species, with subunits Rpo8 and Rpo13 found in restricted subsets of archaea. While Rpo8 has an ortholog, Rpb8, in the nuclear eukaryal RNAPs, Rpo13 lacks clear eukaryal orthologs. Here, we report crystal structures of the DNA-bound and free form of the aRNAP from Sulfolobus shibatae. Together with biochemical and biophysical analyses, these data show that Rpo13 C-terminus binds non-specifically to double-stranded DNA. These interactions map on our RNAP-DNA binary complex on the downstream DNA at the far end of the DNA entry channel. Our findings thus support Rpo13 as a RNAP-DNA stabilization factor, a role reminiscent of eukaryotic general transcriptional factors. The data further yield insight into the mechanisms and evolution of RNAP-DNA interaction.
- Structural Biology Unit, CIC bioGUNE, CIBERehd, 48160 Derio, Spain.
Organizational Affiliation: