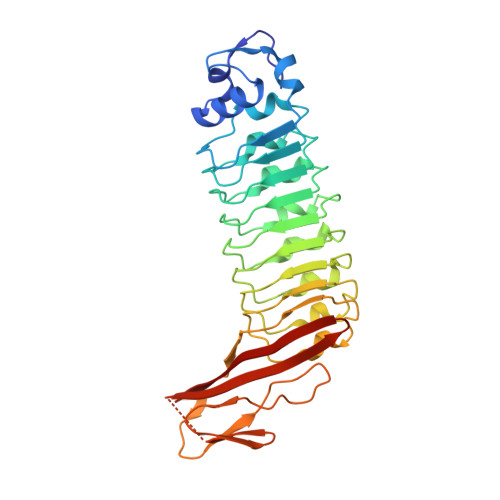
Engineered Variants of Inlb with an Additional Leucine-Rich Repeat Discriminate between Physiologically Relevant and Packing Contacts in Crystal Structures of the Inlb:Met Complex.
Niemann, H.H., Gherardi, E., Bleymuller, W.M., Heinz, D.W.(2012) Protein Sci 21: 1528
- PubMed: 22887347 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2142
- Primary Citation Related Structures:
4AW4 - PubMed Abstract:
The physiological relevance of contacts in crystal lattices often remains elusive. This was also the case for the complex between the invasion protein internalin B (InlB) from Listeria monocytogenes and its host cell receptor, the human receptor tyrosine kinase (RTK) MET. InlB is a MET agonist and induces bacterial host cell invasion. Activation of RTKs generally involves ligand-induced dimerization of the receptor ectodomain. The two currently available crystal structures of the InlB:MET complex show the same arrangement of InlB and MET in a 1:1 complex, but different dimeric 2:2 assemblies. Only one of these 2:2 assemblies is predicted to be stable by a computational procedure. This assembly is mainly stabilized by a contact between the Cap domain of InlB from one and the Sema domain of MET from another 1:1 complex. Here, we probe the physiological relevance of this interaction. We generated variants of the leucine-rich repeat (LRR) protein InlB by inserting an additional repeat between the first and the second LRR. This should allow formation of the 1:1 complex but disrupt the potential 2:2 complex involving the Cap-Sema contact due to steric distortions. A crystal structure of one of the engineered proteins showed that it folded properly. Binding affinity to MET was comparable to that of wild-type InlB. The InlB variant induced MET phosphorylation and cell scatter like wild-type InlB. These results suggest that the Cap-Sema interaction is not physiologically relevant and support the previously proposed assembly, in which a 2:2 InlB:MET complex is built around a ligand dimer.
- Department of Chemistry, Bielefeld University, 33501 Bielefeld, Germany. Hartmut.Niemann@uni-bielefeld.de
Organizational Affiliation: