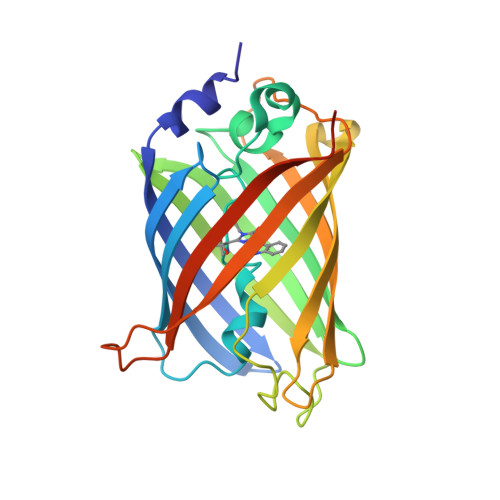
Alteration of Fluorescent Protein Spectroscopic Properties Upon Cryoprotection
von Stetten, D., Batot, G., Noirclerc-Savoye, M., Royant, A.(2012) Acta Crystallogr D Biol Crystallogr 68: 1578
- PubMed: 23090407 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912037900
- Primary Citation Related Structures:
4AS8 - PubMed Abstract:
Cryoprotection of a protein crystal by addition of small-molecule compounds may sometimes affect the structure of its active site. The spectroscopic and structural effects of the two cryoprotectants glycerol and ethylene glycol on the cyan fluorescent protein Cerulean were investigated. While glycerol had almost no noticeable effect, ethylene glycol was shown to induce a systematic red shift of the UV-vis absorption and fluorescence emission spectra. Additionally, ethylene glycol molecules were shown to enter the core of the protein, with one of them binding in close vicinity to the chromophore, which provides a sound explanation for the observed spectroscopic changes. These results highlight the need to systematically record spectroscopic data on crystals of light-absorbing proteins and reinforce the notion that fluorescent proteins must not been seen as rigid structures.
- Structural Biology Group, European Synchrotron Radiation Facility, Grenoble, France. vonstett@esrf.fr
Organizational Affiliation: