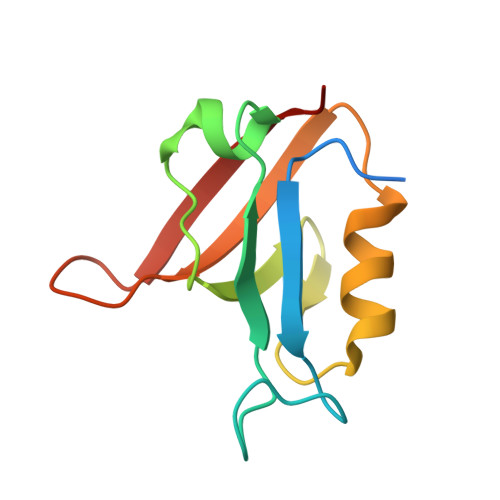
Tolerance of Protein Folding to a Circular Permutation in a Pdz Domain
Hultqvist, G., Punekar, A.S., Morrone, A., Chi, C.N., Engstrom, A., Selmer, M., Gianni, S., Jemth, P.(2012) PLoS One 7: 50055
- PubMed: 23185531 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0050055
- Primary Citation Related Structures:
4AMH - PubMed Abstract:
Circular permutation is a common molecular mechanism for evolution of proteins. However, such re-arrangement of secondary structure connectivity may interfere with the folding mechanism causing accumulation of folding intermediates, which in turn can lead to misfolding. We solved the crystal structure and investigated the folding pathway of a circularly permuted variant of a PDZ domain, SAP97 PDZ2. Our data illustrate how well circular permutation may work as a mechanism for molecular evolution. The circular permutant retains the overall structure and function of the native protein domain. Further, unlike most examples in the literature, this circular permutant displays a folding mechanism that is virtually identical to that of the wild type. This observation contrasts with previous data on the circularly permuted PDZ2 domain from PTP-BL, for which the folding pathway was remarkably affected by the same mutation in sequence connectivity. The different effects of this circular permutation in two homologous proteins show the strong influence of sequence as compared to topology. Circular permutation, when peripheral to the major folding nucleus, may have little effect on folding pathways and could explain why, despite the dramatic change in primary structure, it is frequently tolerated by different protein folds.
- Department of Medical Biochemistry and Microbiology, Uppsala University, Uppsala, Sweden.
Organizational Affiliation: