Evolution from the Prokaryotic to the Higher Plant Chloroplast Signal Recognition Particle: The Signal Recognition Particle RNA is Conserved in Plastids of a Wide Range of Photosynthetic Organisms.
Trager, C., Rosenblad, M.A., Ziehe, D., Garcia-Petit, C., Schrader, L., Kock, K., Vera Richter, C., Klinkert, B., Narberhaus, F., Herrmann, C., Hofmann, E., Aronsson, H., Schunemann, D.(2012) Plant Cell 24: 4819
- PubMed: 23275580 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1105/tpc.112.102996
- Primary Citation Related Structures:
4AK9 - PubMed Abstract:
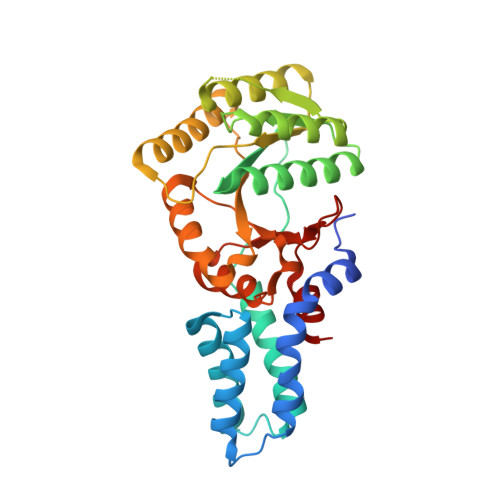
The protein targeting signal recognition particle (SRP) pathway in chloroplasts of higher plants has undergone dramatic evolutionary changes. It disposed of its RNA, which is an essential SRP component in bacteria, and uses a unique chloroplast-specific protein cpSRP43. Nevertheless, homologs of the conserved SRP54 and the SRP receptor, FtsY, are present in higher plant chloroplasts. In this study, we analyzed the phylogenetic distribution of SRP components in photosynthetic organisms to elucidate the evolution of the SRP system. We identified conserved plastid SRP RNAs within all nonspermatophyte land plant lineages and in all chlorophyte branches. Furthermore, we show the simultaneous presence of cpSRP43 in these organisms. The function of this novel SRP system was biochemically and structurally characterized in the moss Physcomitrella patens. We show that P. patens chloroplast SRP (cpSRP) RNA binds cpSRP54 but has lost the ability to significantly stimulate the GTPase cycle of SRP54 and FtsY. Furthermore, the crystal structure at 1.8-Å resolution and the nucleotide specificity of P. patens cpFtsY was determined and compared with bacterial FtsY and higher plant chloroplast FtsY. Our data lead to the view that the P. patens cpSRP system occupies an intermediate position in the evolution from bacterial-type SRP to higher plant-type cpSRP system.
- Molecular Biology of Plant Organelles, Ruhr-University Bochum, 44780 Bochum, Germany.
Organizational Affiliation: