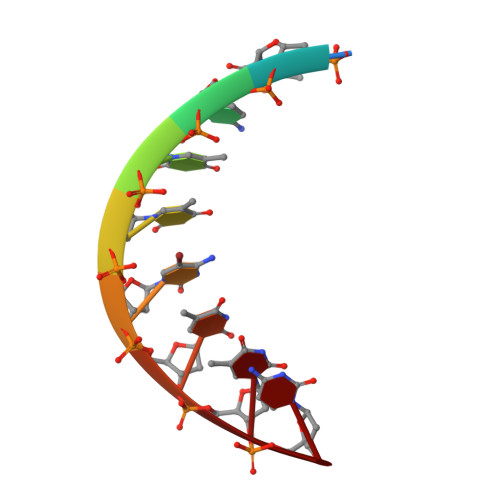
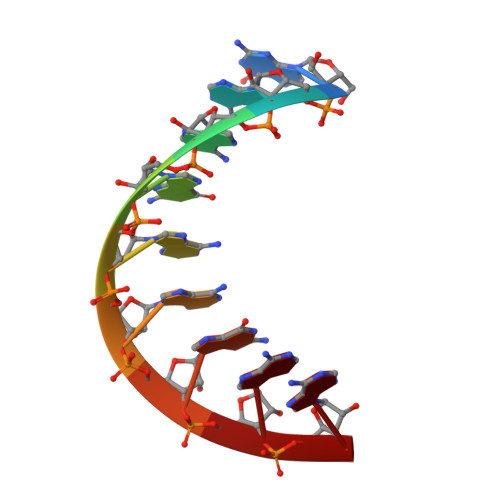
Crystal structure and conformation of a DNA-RNA hybrid duplex with a polypurine RNA strand: d(TTCTTBr5CTTC)-r(GAAGAAGAA).
Xiong, Y., Sundaralingam, M.(1998) Structure 6: 1493-1501
- PubMed: 9862803 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(98)00148-8
- Primary Citation Related Structures:
421D - PubMed Abstract:
. DNA-RNA hybrids are substrates for RNase H. This enzyme catalyzes the hydrolysis of the RNA strand in the hybrid form. The polypurine tract (PPT) in human immunodeficiency virus 1 (HIV-1) is a short stretch of purines ( approximately 15 bases) located at the 3'-end of the U3 region of the RNA genome. The PPT has the unique ability to resist digestion by RNase H and serves as a primer for plus-strand DNA synthesis. . The crystal structure of a DNA-RNA hybrid duplex containing a polypurine RNA strand, d(TTCTTBr5CTTC)-r(GAAGAAGAA), has been determined at 1.8 A resolution. The structure was solved by molecular replacement methods and refined to a final R factor of 20.1% (R free 23.7%). The hybrid duplex adopts a standard A-form conformation. All of the sugar rings and glycosidic torsion angles are found in the standard C3'-endo/anti conformation, as seen in A-RNA or A-DNA. The crystal packing is dominated by the DNA strand, where the terminal base pairs of the hybrid abut the neighboring A-DNA sugar-phosphate backbone on the minor groove side. . The present DNA-RNA hybrid duplex containing a polypurine RNA strand exhibits standard A-form geometry. This observation might suggest that the RNA PPT resists the RNase H activity of HIV reverse transcriptase as a result of its A-form conformation. In addition, there appears to be a correlation between the percentage purine content of the RNA and the DNA backbone conformation.
- The Ohio State University Biological Macromolecular Structure Center Departments of Chemistry, Biochemistry and Biophysics Program 012 Rightmire Hall 1060 Carmack Road Columbus Ohio 43210 USA.
Organizational Affiliation: