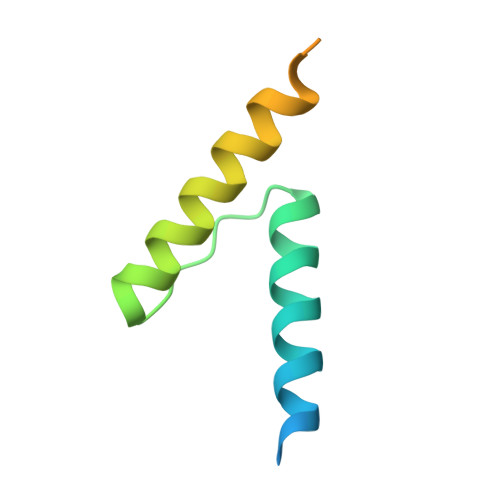
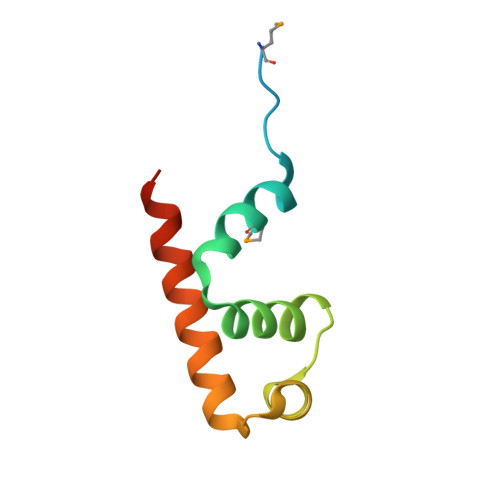
Mycobacterium tuberculosis RsdA provides a conformational rationale for selective regulation of sigma-factor activity by proteolysis
Jaiswal, R.K., Prabha, T.S., Manjeera, G., Gopal, B.(2013) Nucleic Acids Res 41: 3414-3423
- PubMed: 23314154 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gks1468
- Primary Citation Related Structures:
3VEP, 3VFZ - PubMed Abstract:
The relative levels of different σ factors dictate the expression profile of a bacterium. Extracytoplasmic function σ factors synchronize the transcriptional profile with environmental conditions. The cellular concentration of free extracytoplasmic function σ factors is regulated by the localization of this protein in a σ/anti-σ complex. Anti-σ factors are multi-domain proteins with a receptor to sense environmental stimuli and a conserved anti-σ domain (ASD) that binds a σ factor. Here we describe the structure of Mycobacterium tuberculosis anti-σ(D) (RsdA) in complex with the -35 promoter binding domain of σ(D) (σ(D)4). We note distinct conformational features that enable the release of σ(D) by the selective proteolysis of the ASD in RsdA. The structural and biochemical features of the σ(D)/RsdA complex provide a basis to reconcile diverse regulatory mechanisms that govern σ/anti-σ interactions despite high overall structural similarity. Multiple regulatory mechanisms embedded in an ASD scaffold thus provide an elegant route to rapidly re-engineer the expression profile of a bacterium in response to an environmental stimulus.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560 012, India.
Organizational Affiliation: