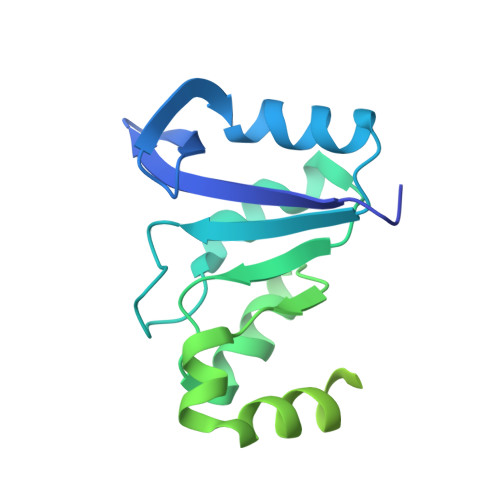
Exploring symmetry as an avenue to the computational design of large protein domains.
Fortenberry, C., Bowman, E.A., Proffitt, W., Dorr, B., Combs, S., Harp, J., Mizoue, L., Meiler, J.(2011) J Am Chem Soc 133: 18026-18029
- PubMed: 21978247 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja2051217
- Primary Citation Related Structures:
3TDM, 3TDN - PubMed Abstract:
It has been demonstrated previously that symmetric, homodimeric proteins are energetically favored, which explains their abundance in nature. It has been proposed that such symmetric homodimers underwent gene duplication and fusion to evolve into protein topologies that have a symmetric arrangement of secondary structure elements--"symmetric superfolds". Here, the ROSETTA protein design software was used to computationally engineer a perfectly symmetric variant of imidazole glycerol phosphate synthase and its corresponding symmetric homodimer. The new protein, termed FLR, adopts the symmetric (βα)(8) TIM-barrel superfold. The protein is soluble and monomeric and exhibits two-fold symmetry not only in the arrangement of secondary structure elements but also in sequence and at atomic detail, as verified by crystallography. When cut in half, FLR dimerizes readily to form the symmetric homodimer. The successful computational design of FLR demonstrates progress in our understanding of the underlying principles of protein stability and presents an attractive strategy for the in silico construction of larger protein domains from smaller pieces.
- Department of Chemistry, Vanderbilt University, Nashville, Tennessee 37235, United States.
Organizational Affiliation: