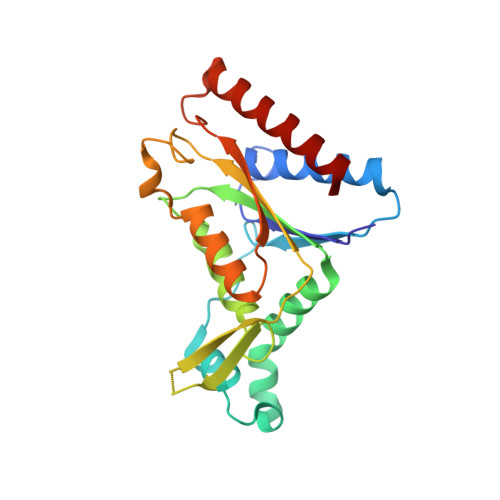
Activation of nitrofurazone by azoreductases: multiple activities in one enzyme.
Ryan, A., Kaplan, E., Laurieri, N., Lowe, E., Sim, E.(2011) Sci Rep 1: 63-63
- PubMed: 22355582 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep00063
- Primary Citation Related Structures:
3R6W - PubMed Abstract:
Azoreductases are well known for azo pro-drug activation by gut flora. We show that azoreductases have a wider role in drug metabolism than previously thought as they can also reduce and hence activate nitrofurazone. Nitrofurazone, a nitroaromatic drug, is a broad spectrum antibiotic which has until now been considered as activated in bacteria by nitroreductases. The structure of the azoreductase with nitrofurazone bound was solved at 2.08 Å and shows nitrofurazone in an active conformation. Based on the structural information, the kinetics and stoichiometry of nitrofurazone reduction by azoreductase from P. aeruginosa, we propose a mechanism of activation which accounts for the ability of azoreductases to reduce both azo and nitroaromatic drugs. This mode of activation can explain the cytotoxic side-effects of nitrofurazone through human azoreductase homologues.
- Pharmacology Department, University of Oxford, OX1 3QT.
Organizational Affiliation: