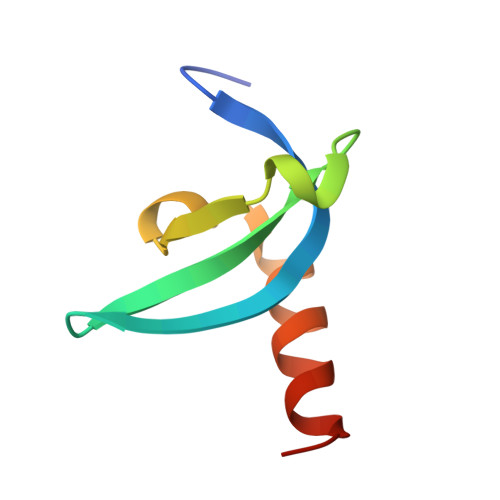
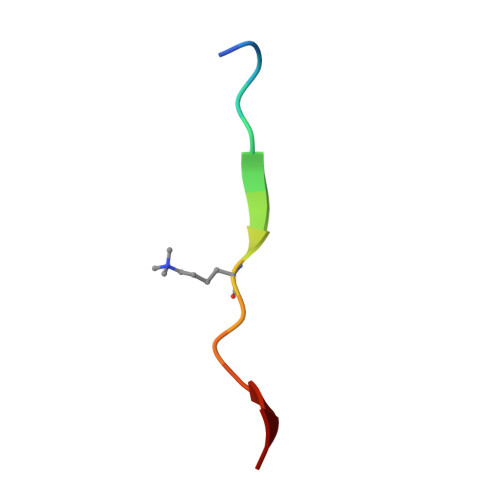
Structural insights for MPP8 chromodomain interaction with histone H3 lysine 9: potential effect of phosphorylation on methyl-lysine binding.
Chang, Y., Horton, J.R., Bedford, M.T., Zhang, X., Cheng, X.(2011) J Mol Biology 408: 807-814
- PubMed: 21419134 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.03.018
- Primary Citation Related Structures:
3QO2 - PubMed Abstract:
M-phase phosphoprotein 8 (MPP8) harbors an N-terminal chromodomain and a C-terminal ankyrin repeat domain. MPP8, via its chromodomain, binds histone H3 peptide tri- or di-methylated at lysine 9 (H3K9me3/H3K9me2) in submicromolar affinity. We determined the crystal structure of MPP8 chromodomain in complex with H3K9me3 peptide. MPP8 interacts with at least six histone H3 residues from glutamine 5 to serine 10, enabling its ability to distinguish lysine-9-containing peptide (QTARKS) from that of lysine 27 (KAARKS), both sharing the ARKS sequence. A partial hydrophobic cage with three aromatic residues (Phe59, Trp80 and Tyr83) and one aspartate (Asp87) encloses the methylated lysine 9. MPP8 has been reported to be phosphorylated in vivo, including the cage residue Tyr83 and the succeeding Thr84 and Ser85. Modeling a phosphate group onto the side-chain hydroxyl oxygen of Tyr83 suggests that the negatively charged phosphate group could enhance the binding of positively charged methyl-lysine or create a regulatory signal by allowing or inhibiting binding of other protein(s).
- Department of Biochemistry, Emory University School of Medicine, Atlanta, GA 30322, USA.
Organizational Affiliation: