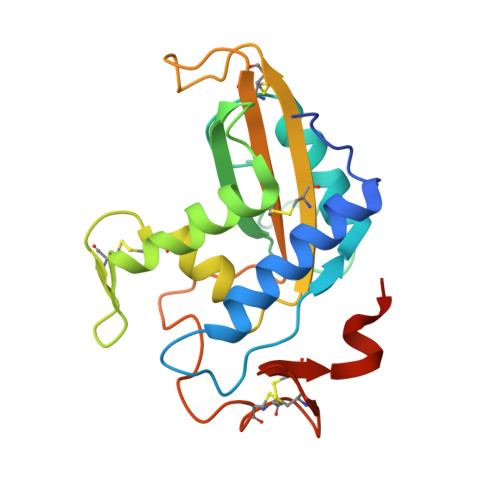
Structural studies of human glioma pathogenesis-related protein 1.
Asojo, O.A., Koski, R.A., Bonafe, N.(2011) Acta Crystallogr D Biol Crystallogr 67: 847-855
- PubMed: 21931216 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444911028198
- Primary Citation Related Structures:
3Q2R, 3Q2U - PubMed Abstract:
Human glioma pathogenesis-related protein 1 (GLIPR1) is a membrane protein that is highly upregulated in brain cancers but is barely detectable in normal brain tissue. GLIPR1 is composed of a signal peptide that directs its secretion, a conserved cysteine-rich CAP (cysteine-rich secretory proteins, antigen 5 and pathogenesis-related 1 proteins) domain and a transmembrane domain. GLIPR1 is currently being investigated as a candidate for prostate cancer gene therapy and for glioblastoma targeted therapy. Crystal structures of a truncated soluble domain of the human GLIPR1 protein (sGLIPR1) solved by molecular replacement using a truncated polyalanine search model of the CAP domain of stecrisp, a snake-venom cysteine-rich secretory protein (CRISP), are presented. The correct molecular-replacement solution could only be obtained by removing all loops from the search model. The native structure was refined to 1.85 Å resolution and that of a Zn2+ complex was refined to 2.2 Å resolution. The latter structure revealed that the putative binding cavity coordinates Zn2+ similarly to snake-venom CRISPs, which are involved in Zn2+-dependent mechanisms of inflammatory modulation. Both sGLIPR1 structures have extensive flexible loop/turn regions and unique charge distributions that were not observed in any of the previously reported CAP protein structures. A model is also proposed for the structure of full-length membrane-bound GLIPR1.
- Department of Pathology and Microbiology, College of Medicine, Nebraska Medical Center, Omaha, NE 68198-6495, USA. oasojo@unmc.edu
Organizational Affiliation: