Structural Basis of Streptococcus pyogenes Immunity to Its NAD(+) Glycohydrolase Toxin.
Smith, C.L., Ghosh, J., Elam, J.S., Pinkner, J.S., Hultgren, S.J., Caparon, M.G., Ellenberger, T.(2011) Structure 19: 192-202
- PubMed: 21300288 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2010.12.013
- Primary Citation Related Structures:
3PNT, 3QB2 - PubMed Abstract:
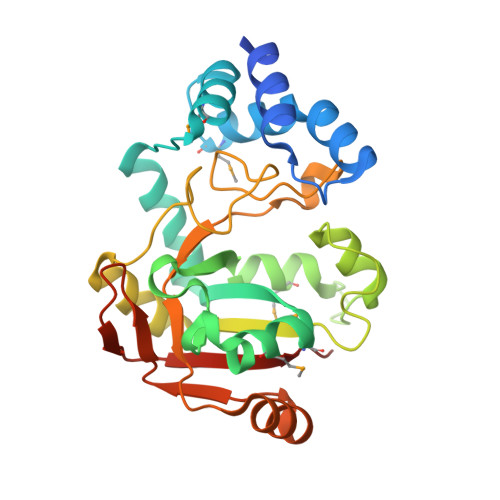
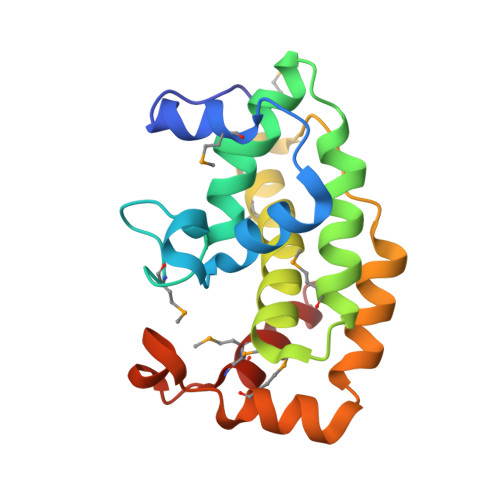
The virulence of Gram-positive bacteria is enhanced by toxins like the Streptococcus pyogenes β-NAD(+) glycohydrolase known as SPN. SPN-producing strains of S. pyogenes additionally express the protein immunity factor for SPN (IFS), which forms an inhibitory complex with SPN. We have determined crystal structures of the SPN-IFS complex and IFS alone, revealing that SPN is structurally related to ADP-ribosyl transferases but lacks the canonical binding site for protein substrates. SPN is instead a highly efficient glycohydrolase with the potential to deplete cellular levels of β-NAD(+). The protective effect of IFS involves an extensive interaction with the SPN active site that blocks access to β-NAD(+). The conformation of IFS changes upon binding to SPN, with repacking of an extended C-terminal α helix into a compact shape. IFS is an attractive target for the development of novel bacteriocidal compounds functioning by blocking the bacterium's self-immunity to the SPN toxin.
- Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, Saint Louis, MO 63110-1093, USA.
Organizational Affiliation: