Crystal Structure of Acyl-CoA Dehydrogenase complexed with FAD from Bacillus anthracis
Kim, Y., Maltseva, N., Kwon, K., Anderson, W.F., Joachimiak, A., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Acyl-CoA dehydrogenase | 597 | Bacillus anthracis str. 'Ames Ancestor | Mutation(s): 0 Gene Names: BAS4875, BA_5246, GBAA5246, GBAA_5246 EC: 1.3.99 | 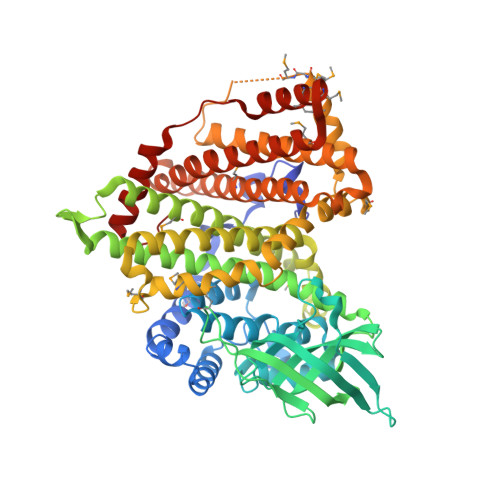 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A6H3ADM7 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| FAD Download:Ideal Coordinates CCD File | E [auth A], J [auth B], R [auth C], Y [auth D] | FLAVIN-ADENINE DINUCLEOTIDE C27 H33 N9 O15 P2 VWWQXMAJTJZDQX-UYBVJOGSSA-N |  | ||
| 1PE Download:Ideal Coordinates CCD File | F [auth A], M [auth B], U [auth C], Z [auth D] | PENTAETHYLENE GLYCOL C10 H22 O6 JLFNLZLINWHATN-UHFFFAOYSA-N |  | ||
| PEG Download:Ideal Coordinates CCD File | G [auth A], N [auth B], W [auth C], X [auth D] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| SO4 Download:Ideal Coordinates CCD File | I [auth A], O [auth B] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| GOL Download:Ideal Coordinates CCD File | H [auth A] K [auth B] L [auth B] P [auth B] Q [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MSE Query on MSE | A, B, C, D | L-PEPTIDE LINKING | C5 H11 N O2 Se |  | MET |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 75.613 | α = 92.8 |
| b = 98.028 | β = 106.63 |
| c = 107.743 | γ = 105.3 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| HKL-3000 | data collection |
| BALBES | phasing |
| PHENIX | refinement |
| SBC-Collect | data collection |
| HKL-3000 | data reduction |
| HKL-3000 | data scaling |