Selection of a novel and highly specific TNF{alpha} antagonist: insight from the crystal structure of the antagonist-TNF{alpha} complex
Byla, P., Andersen, M.H., Holtet, T.L., Jacobsen, H., Munch, M., Gad, H.H., Thogersen, H.C., Hartmann, R.(2010) J Biological Chem 285: 12096-12100
- PubMed: 20179326 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.063305
- Primary Citation Related Structures:
3L9J - PubMed Abstract:
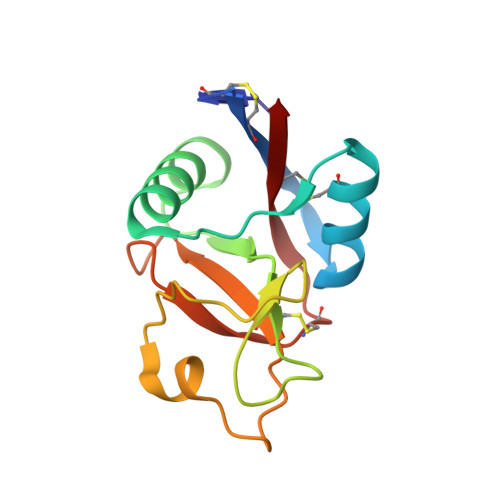
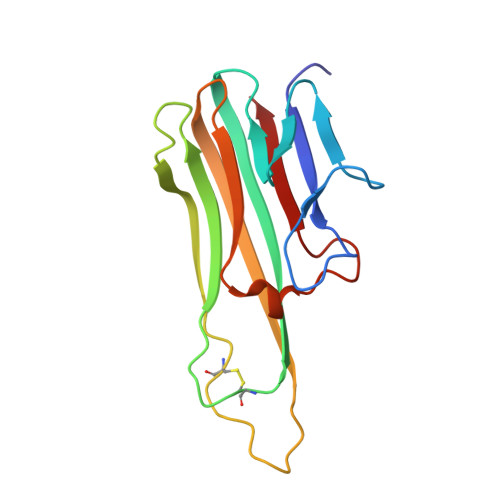
Inhibition of tumor necrosis factor alpha (TNFalpha) is a favorable way of treating several important diseases such as rheumatoid arthritis, Crohn disease, and psoriasis. Therefore, an extensive range of TNFalpha inhibitory proteins, most of them based upon an antibody scaffold, has been developed and used with variable success as therapeutics. We have developed a novel technology platform using C-type lectins as a vehicle for the creation of novel trimeric therapeutic proteins with increased avidity and unique properties as compared with current protein therapeutics. We chose human TNFalpha as a test target to validate this new technology because of the extensive experience available with protein-based TNFalpha antagonists. Here, we present a novel and highly specific TNFalpha antagonist developed using this technology. Furthermore, we have solved the three-dimensional structure of the antagonist-TNFalpha complex by x-ray crystallography, and this structure is presented here. The structure has given us a unique insight into how the selection procedure works at a molecular level. Surprisingly little change is observed in the C-type lectin-like domain structure outside of the randomized regions, whereas a substantial change is observed within the randomized loops. Thus, the overall integrity of the C-type lectin-like domain is maintained, whereas specificity and binding affinity are changed by the introduction of a number of specific contacts with TNFalpha.
- Centre for Structural Biology, Department of Molecular Biology, Aarhus University, Gustav Wieds Vej 10C, 8000 Aarhus C, Denmark.
Organizational Affiliation: