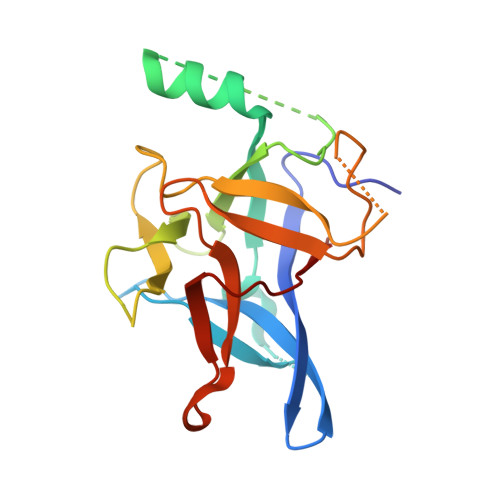
Crystal structures of the N-terminal domains of cardiac and skeletal muscle ryanodine receptors: insights into disease mutations.
Lobo, P.A., Van Petegem, F.(2009) Structure 17: 1505-1514
- PubMed: 19913485 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2009.08.016
- Primary Citation Related Structures:
3ILA, 3IM5, 3IM6, 3IM7 - PubMed Abstract:
Ryanodine receptors (RyRs) are channels governing the release of Ca(2+) from the sarcoplasmic or endoplasmic reticulum. They are required for the contraction of both skeletal (RyR1) and cardiac (RyR2) muscles. Mutations in both RyR1 and RyR2 have been associated with severe genetic disorders, but high-resolution data describing the disease variants in detail have been lacking. Here we present the crystal structures of the N-terminal domains of both RyR2 (1-217) and RyR1 (9-205) at 2.55 A and 2.9 A, respectively. The domains map in a hot spot region for disease mutations. Both structures consist of a core beta trefoil domain flanked by an alpha helix. Crystal structures of two RyR2 disease mutants, A77V (2.2 A) and V186M (1.7 A), show that the mutations cause distinct local changes in the surface of the protein. A RyR2 deletion mutant causes significant changes in the thermal stability. The disease positions highlight two putative binding interfaces required for normal RyR function.
- University of British Columbia, Department of Biochemistry and Molecular Biology, Vancouver, BC V6T 1Z3 Canada.
Organizational Affiliation: