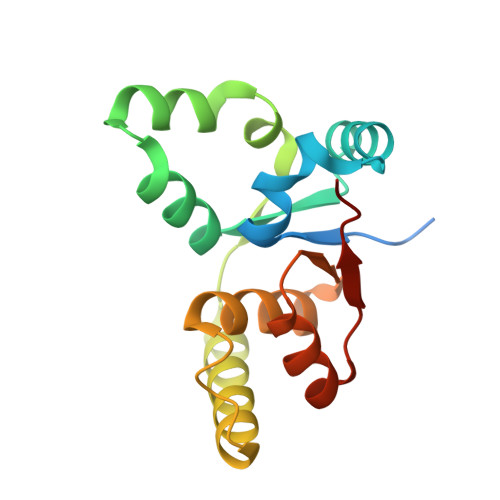
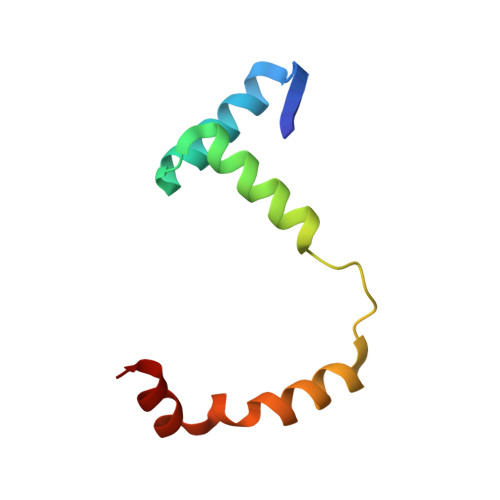
The crystal structure of the Rv0301-Rv0300 VapBC-3 toxin-antitoxin complex from M. tuberculosis reveals a Mg(2+) ion in the active site and a putative RNA-binding site.
Min, A.B., Miallau, L., Sawaya, M.R., Habel, J., Cascio, D., Eisenberg, D.(2012) Protein Sci 21: 1754-1767
- PubMed: 23011806 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2161
- Primary Citation Related Structures:
3H87 - PubMed Abstract:
VapBC pairs account for 45 out of 88 identified toxin-antitoxin (TA) pairs in the Mycobacterium tuberculosis (Mtb) H37Rv genome. A working model suggests that under times of stress, antitoxin molecules are degraded, releasing the toxins to slow the metabolism of the cell, which in the case of VapC toxins is via their RNase activity. Otherwise the TA pairs remain bound to their promoters, autoinhibiting transcription. The crystal structure of Rv0301-Rv0300, an Mtb VapBC TA complex determined at 1.49 Å resolution, suggests a mechanism for these three functions: RNase activity, its inhibition by antitoxin, and its ability to bind promoter DNA. The Rv0301 toxin consists of a core of five parallel beta strands flanked by alpha helices. Three proximal aspartates coordinate a Mg²⁺ ion forming the putative RNase active site. The Rv0300 antitoxin monomer is extended in structure, consisting of an N-terminal beta strand followed by four helices. The last two helices wrap around the toxin and terminate near the putative RNase active site, but with different conformations. In one conformation, the C-terminal arginine interferes with Mg²⁺ ion coordination, suggesting a mechanism by which the antitoxin can inhibit toxin activity. At the N-terminus of the antitoxin, two pairs of Ribbon-Helix-Helix (RHH) motifs are related by crystallographic twofold symmetry. The resulting hetero-octameric complex is similar to the FitAB system, but the two RHH motifs are about 30 Å closer together in the Rv0301-Rv0300 complex, suggesting either a different span of the DNA recognition sequence or a conformational change.
- Department of Chemistry and Biochemistry, Howard Hughes Medical Institute, UCLA-DOE Institute of Genomics and Proteomics, UCLA, Los Angeles, California 90095-1570, USA.
Organizational Affiliation: