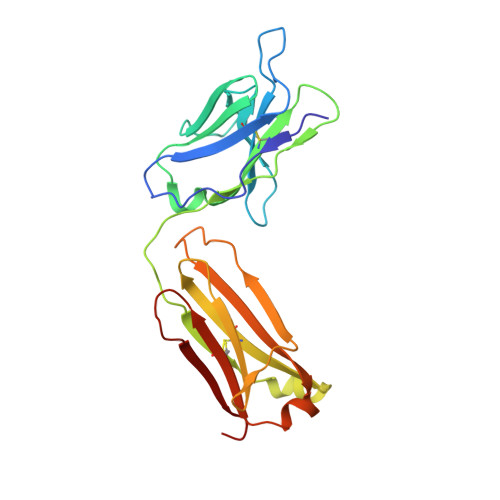
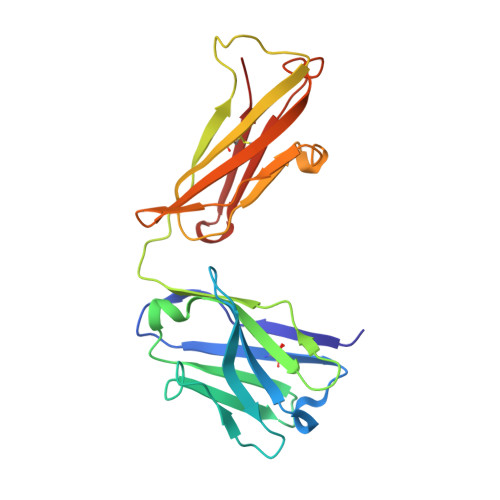
Deeply inverted electron-hole recombination in a luminescent antibody-stilbene complex.
Debler, E.W., Kaufmann, G.F., Meijler, M.M., Heine, A., Mee, J.M., Pljevaljcic, G., Di Bilio, A.J., Schultz, P.G., Millar, D.P., Janda, K.D., Wilson, I.A., Gray, H.B., Lerner, R.A.(2008) Science 319: 1232-1235
- PubMed: 18309081 Search on PubMed
- DOI: https://doi.org/10.1126/science.1153445
- Primary Citation Related Structures:
3CFB, 3CFC, 3CFD, 3CFE - PubMed Abstract:
The blue-emissive antibody EP2-19G2 that has been elicited against trans-stilbene has unprecedented ability to produce bright luminescence and has been used as a biosensor in various applications. We show that the prolonged luminescence is not stilbene fluorescence. Instead, the emissive species is a charge-transfer excited complex of an anionic stilbene and a cationic, parallel pi-stacked tryptophan. Upon charge recombination, this complex generates exceptionally bright blue light. Complex formation is enabled by a deeply penetrating ligand-binding pocket, which in turn results from a noncanonical interface between the two variable domains of the antibody.
- Department of Molecular Biology, Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: