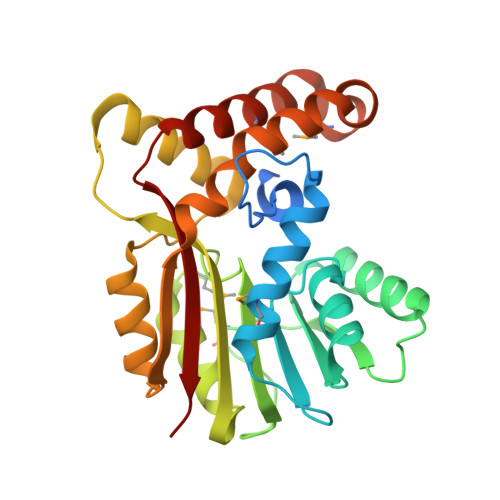
Structure and mechanism of the rebeccamycin sugar 4'-O-methyltransferase RebM.
Singh, S., McCoy, J.G., Zhang, C., Bingman, C.A., Phillips Jr., G.N., Thorson, J.S.(2008) J Biological Chem 283: 22628-22636
- PubMed: 18502766 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M800503200
- Primary Citation Related Structures:
3BUS - PubMed Abstract:
The 2.65-angstroms crystal structure of the rebeccamycin 4'-O-methyltransferase RebM in complex with S-adenosyl-l-homocysteine revealed RebM to adopt a typical S-adenosylmethionine-binding fold of small molecule O-methyltransferases (O-MTases) and display a weak dimerization domain unique to MTases. Using this structure as a basis, the RebM substrate binding model implicated a predominance of nonspecific hydrophobic interactions consistent with the reported ability of RebM to methylate a wide range of indolocarbazole surrogates. This model also illuminated the three putative RebM catalytic residues (His140/141 and Asp166) subsequently found to be highly conserved among sequence-related natural product O-MTases from GC-rich bacteria. Interrogation of these residues via site-directed mutagenesis in RebM demonstrated His140 and Asp166 to be most important for catalysis. This study reveals RebM to be a member of the general acid/base-dependent O-MTases and, as the first crystal structure for a sugar O-MTase, may also present a template toward the future engineering of natural product MTases for combinatorial applications.
- Division of Pharmaceutical Sciences, School of Pharmacy, University of Wisconsin, Madison, Wisconsin 53705, USA.
Organizational Affiliation: