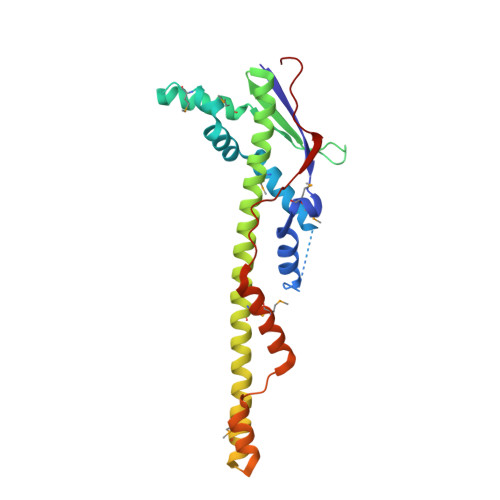
Bacterial polysaccharide co-polymerases share a common framework for control of polymer length
Tocilj, A., Munger, C., Proteau, A., Morona, R., Purins, L., Ajamian, E., Wagner, J., Papadopoulos, M., Van Den Bosch, L., Rubinstein, J.L., Fethiere, J., Matte, A., Cygler, M.(2008) Nat Struct Mol Biol 15: 130-138
- PubMed: 18204465 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.1374
- Primary Citation Related Structures:
3B8M, 3B8N, 3B8O, 3B8P - PubMed Abstract:
The chain length distribution of complex polysaccharides present on the bacterial surface is determined by polysaccharide co-polymerases (PCPs) anchored in the inner membrane. We report crystal structures of the periplasmic domains of three PCPs that impart substantially different chain length distributions to surface polysaccharides. Despite very low sequence similarities, they have a common protomer structure with a long central alpha-helix extending 100 A into the periplasm. The protomers self-assemble into bell-shaped oligomers of variable sizes, with a large internal cavity. Electron microscopy shows that one of the full-length PCPs has a similar organization as that observed in the crystal for its periplasmic domain alone. Functional studies suggest that the top of the PCP oligomers is an important region for determining polysaccharide modal length. These structures provide a detailed view of components of the bacterial polysaccharide assembly machinery.
- Department of Biochemistry, McGill University, 3655 Promenade Sir William Osler, Montréal, Québec H3G 1Y6, Canada.
Organizational Affiliation: