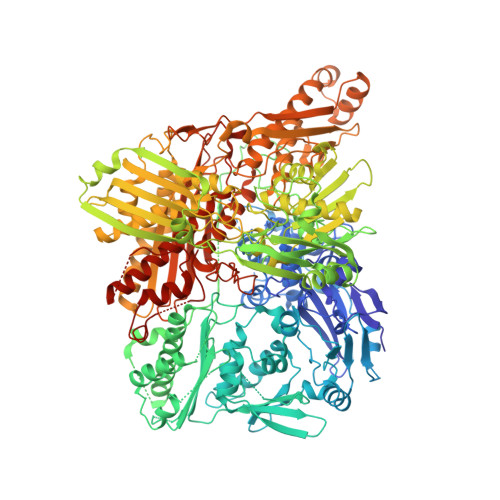
The First Mammalian Aldehyde Oxidase Crystal Structure: Insights Into Substrate Specificity.
Coelho, C., Mahro, M., Trincao, J., Carvalho, A.T.P., Ramos, M.J., Terao, M., Garattini, E., Leimkuhler, S., Romao, M.J.(2012) J Biological Chem 287: 40690
- PubMed: 23019336 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.390419
- Primary Citation Related Structures:
3ZYV - PubMed Abstract:
Aldehyde oxidases have pharmacological relevance, and AOX3 is the major drug-metabolizing enzyme in rodents. The crystal structure of mouse AOX3 with kinetics and molecular docking studies provides insights into its enzymatic characteristics. Differences in substrate and inhibitor specificities can be rationalized by comparing the AOX3 and xanthine oxidase structures. The first aldehyde oxidase structure represents a major advance for drug design and mechanistic studies. Aldehyde oxidases (AOXs) are homodimeric proteins belonging to the xanthine oxidase family of molybdenum-containing enzymes. Each 150-kDa monomer contains a FAD redox cofactor, two spectroscopically distinct [2Fe-2S] clusters, and a molybdenum cofactor located within the protein active site. AOXs are characterized by broad range substrate specificity, oxidizing different aldehydes and aromatic N-heterocycles. Despite increasing recognition of its role in the metabolism of drugs and xenobiotics, the physiological function of the protein is still largely unknown. We have crystallized and solved the crystal structure of mouse liver aldehyde oxidase 3 to 2.9 Å. This is the first mammalian AOX whose structure has been solved. The structure provides important insights into the protein active center and further evidence on the catalytic differences characterizing AOX and xanthine oxidoreductase. The mouse liver aldehyde oxidase 3 three-dimensional structure combined with kinetic, mutagenesis data, molecular docking, and molecular dynamics studies make a decisive contribution to understand the molecular basis of its rather broad substrate specificity.
- Requimte, Departamento de Química, Faculdade de Ciências e Tecnologia, Universidade Nova de Lisboa, 2829-516 Caparica, Portugal.
Organizational Affiliation: