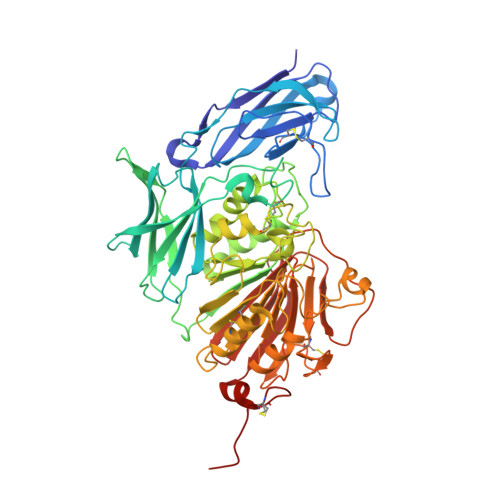
The Structure of a Purple Acid Phosphatase Involved in Plant Growth and Pathogen Defence Exhibits a Novel Immunoglobulin-Like Fold
Antonyuk, S.V., Olczak, M., Olczak, T., Ciuraszkiewicz, J., Strange, R.W.(2014) IUCrJ 1: 101
- PubMed: 25075326 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S205225251400400X
- Primary Citation Related Structures:
3ZK4 - PubMed Abstract:
Phosphatases function in the production, transport and recycling of inorganic phosphorus, which is crucial for cellular metabolism and bioenergetics, as well as in bacterial killing, since they are able to generate reactive oxygen species via Fenton chemistry. Diphosphonucleotide phosphatase/phosphodiesterase (PPD1), a glycoprotein plant purple acid phosphatase (PAP) from yellow lupin seeds, contains a bimetallic Fe-Mn catalytic site which is most active at acidic pH. Unlike other plant PAPs, PPD1 cleaves the pyrophosphate bond in diphosphonucleotides and the phosphodiester bond in various phosphodiesters. The homohexameric organization of PPD1, as revealed by a 1.65 Å resolution crystal structure and confirmed by solution X-ray scattering, is unique among plant PAPs, for which only homodimers have previously been reported. A phosphate anion is bound in a bidentate fashion at the active site, bridging the Fe and Mn atoms in a binding mode similar to that previously reported for sweet potato PAP, which suggests that common features occur in their catalytic mechanisms. The N-terminal domain of PPD1 has an unexpected and unique fibronectin type III-like fold that is absent in other plant PAPs. Here, the in vitro DNA-cleavage activity of PPD1 is demonstrated and it is proposed that the fibronectin III-like domain, which 'overhangs' the active site, is involved in DNA selectivity, binding and activation. The degradation of DNA by PPD1 implies a role for PPD1 in plant growth and repair and in pathogen defence.
- Molecular Biophysics Group, Faculty of Health and Life Sciences, University of Liverpool , Crown Street, Liverpool L69 7ZB, England.
Organizational Affiliation: