How Allosteric Control of Staphylococcus Aureus Penicillin Binding Protein 2A Enables Methicillin Resistance and Physiological Function
Otero, L.H., Rojas-Altuve, A., Llarrull, L.I., Carrasco-Lopez, C., Kumarasiri, M., Lastochkin, E., Fishovitz, J., Dawley, M., Hesek, D., Lee, M., Johnson, J.W., Fisher, J.F., Chang, M., Mobashery, S., Hermoso, J.A.(2013) Proc Natl Acad Sci U S A 110: 16808
- PubMed: 24085846 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1300118110
- Primary Citation Related Structures:
3ZFZ, 3ZG0, 3ZG5 - PubMed Abstract:
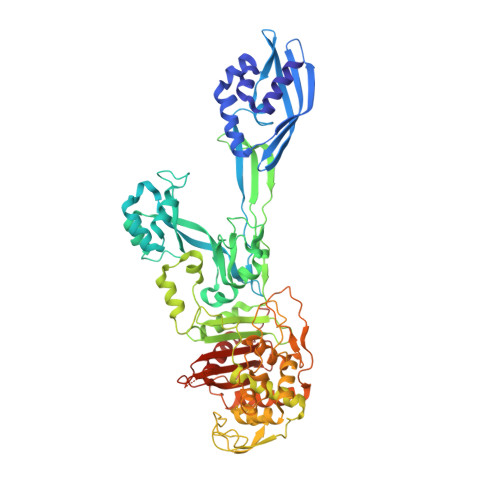
The expression of penicillin binding protein 2a (PBP2a) is the basis for the broad clinical resistance to the β-lactam antibiotics by methicillin-resistant Staphylococcus aureus (MRSA). The high-molecular mass penicillin binding proteins of bacteria catalyze in separate domains the transglycosylase and transpeptidase activities required for the biosynthesis of the peptidoglycan polymer that comprises the bacterial cell wall. In bacteria susceptible to β-lactam antibiotics, the transpeptidase activity of their penicillin binding proteins (PBPs) is lost as a result of irreversible acylation of an active site serine by the β-lactam antibiotics. In contrast, the PBP2a of MRSA is resistant to β-lactam acylation and successfully catalyzes the DD-transpeptidation reaction necessary to complete the cell wall. The inability to contain MRSA infection with β-lactam antibiotics is a continuing public health concern. We report herein the identification of an allosteric binding domain--a remarkable 60 Å distant from the DD-transpeptidase active site--discovered by crystallographic analysis of a soluble construct of PBP2a. When this allosteric site is occupied, a multiresidue conformational change culminates in the opening of the active site to permit substrate entry. This same crystallographic analysis also reveals the identity of three allosteric ligands: muramic acid (a saccharide component of the peptidoglycan), the cell wall peptidoglycan, and ceftaroline, a recently approved anti-MRSA β-lactam antibiotic. The ability of an anti-MRSA β-lactam antibiotic to stimulate allosteric opening of the active site, thus predisposing PBP2a to inactivation by a second β-lactam molecule, opens an unprecedented realm for β-lactam antibiotic structure-based design.
- Departamento de Cristalografía y Biología Estructural, Instituto de Química-Física "Rocasolano," Consejo Superior de Investigaciones Cientificas, 28006 Madrid, Spain.
Organizational Affiliation: