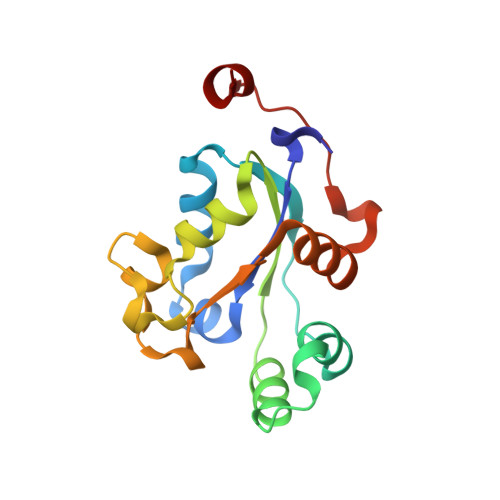
Purification, characterization and structure of nucleoside diphosphate kinase from Drosophila melanogaster
Qian, L., Liu, X.(2014) Protein Expr Purif 103: 48-55
- PubMed: 25195176 Search on PubMed
- DOI: https://doi.org/10.1016/j.pep.2014.08.014
- Primary Citation Related Structures:
3WX8 - PubMed Abstract:
Nucleoside diphosphate kinase (NDPK) is a ubiquitous enzyme found in all organisms and cell types, which catalyzes the transfer of the phosphoryl group from a nucleoside triphosphate to a nucleoside diphosphate. The gene encoding for NDPK from Drosophila melanogaster was amplified from the genomic DNA. The recombinant NDPK (rNDPK) was overexpressed in Escherichia coli and purified to homogeneity by Ni-NTA agarose affinity chromatography, HiTrap SP HP cation exchange chromatography and HiLoad 16/60 Superdex 200 gel filtration chromatography. The gel filtration chromatography and analytical ultracentrifugation showed that rNDPK was a trimer in solution. The binding affinity of NDPs with rNDPK, measured by isothermal titration calorimetry, indicated that the purines nucleotides show higher binding affinity compared with pyrimidines. The rNDPK had a definite nuclease activity in vitro, which could cleave supercoiled plasmid DNA, but had no effect on dsDNA and ssDNA. Furthermore, the structure for NDPK was determined by using the sitting drop vapor diffusion method. In the final model, the asymmetric unit is made of three molecules, each of which consists of a four-stranded anti-parallel β-sheets and seven α-helices. Sequence alignment and structure comparison illustrated that the simulated nucleotide-binding active site are conserved.
- State Key Laboratory of Medicinal Chemical Biology, College of Life Sciences, Nankai University, Tianjin 300071, China.
Organizational Affiliation: