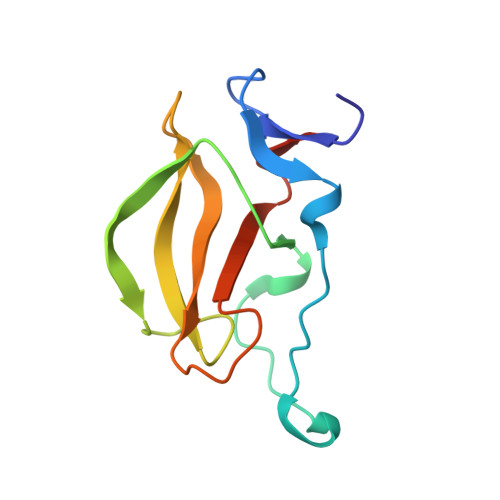
Structure of the human Cereblon-DDB1-lenalidomide complex reveals basis for responsiveness to thalidomide analogs
Chamberlain, P.P., Lopez-Girona, A., Miller, K., Carmel, G., Pagarigan, B., Chie-Leon, B., Rychak, E., Corral, L.G., Ren, Y.J., Wang, M., Riley, M., Delker, S.L., Ito, T., Ando, H., Mori, T., Hirano, Y., Handa, H., Hakoshima, T., Daniel, T.O., Cathers, B.E.(2014) Nat Struct Mol Biol 21: 803-809
- PubMed: 25108355 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2874
- Primary Citation Related Structures:
3WX1, 3WX2 - PubMed Abstract:
The Cul4-Rbx1-DDB1-Cereblon E3 ubiquitin ligase complex is the target of thalidomide, lenalidomide and pomalidomide, therapeutically important drugs for multiple myeloma and other B-cell malignancies. These drugs directly bind Cereblon (CRBN) and promote the recruitment of substrates Ikaros (IKZF1) and Aiolos (IKZF3) to the E3 complex, thus leading to substrate ubiquitination and degradation. Here we present the crystal structure of human CRBN bound to DDB1 and the drug lenalidomide. A hydrophobic pocket in the thalidomide-binding domain (TBD) of CRBN accommodates the glutarimide moiety of lenalidomide, whereas the isoindolinone ring is exposed to solvent. We also solved the structures of the mouse TBD in the apo state and with thalidomide or pomalidomide. Site-directed mutagenesis in lentiviral-expression myeloma models showed that key drug-binding residues are critical for antiproliferative effects.
- Celgene Corporation, San Diego, California, USA.
Organizational Affiliation: