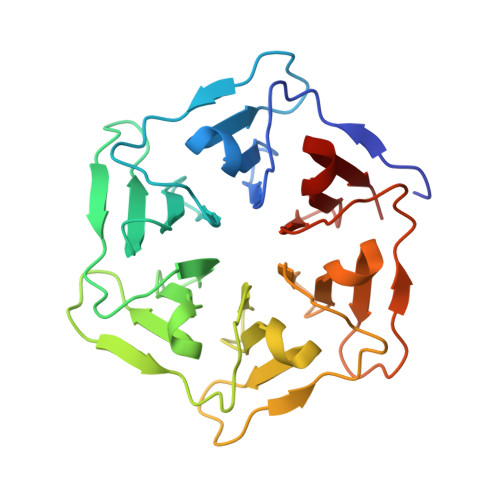
Computational design of a self-assembling symmetrical beta-propeller protein.
Voet, A.R.D., Noguchi, H., Addy, C., Simoncini, D., Terada, D., Unzai, S., Park, S.Y., Zhang, K.Y.J., Tame, J.R.H.(2014) Proc Natl Acad Sci U S A 111: 15102-15107
- PubMed: 25288768 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1412768111
- Primary Citation Related Structures:
3WW7, 3WW8, 3WW9, 3WWA, 3WWB, 3WWF - PubMed Abstract:
The modular structure of many protein families, such as β-propeller proteins, strongly implies that duplication played an important role in their evolution, leading to highly symmetrical intermediate forms. Previous attempts to create perfectly symmetrical propeller proteins have failed, however. We have therefore developed a new and rapid computational approach to design such proteins. As a test case, we have created a sixfold symmetrical β-propeller protein and experimentally validated the structure using X-ray crystallography. Each blade consists of 42 residues. Proteins carrying 2-10 identical blades were also expressed and purified. Two or three tandem blades assemble to recreate the highly stable sixfold symmetrical architecture, consistent with the duplication and fusion theory. The other proteins produce different monodisperse complexes, up to 42 blades (180 kDa) in size, which self-assemble according to simple symmetry rules. Our procedure is suitable for creating nano-building blocks from different protein templates of desired symmetry.
- Structural Bioinformatics Team, Division of Structural and Synthetic Biology, Center for Life Science Technologies, RIKEN, 1-7-22 Suehiro, Yokohama, Kanagawa 230-0045, Japan; and Drug Design Laboratory, Graduate School of Medical Life Science, Yokohama City University, 1-7-29 Suehiro, Yokohama, Kanagawa 230-0045, Japan.
Organizational Affiliation: