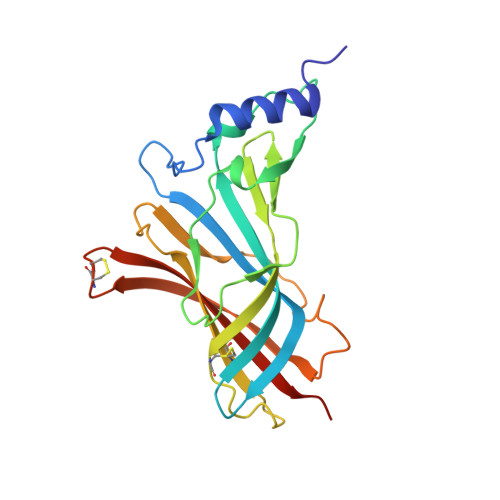
Studies on an acetylcholine binding protein identify a basic residue in loop G on the beta 1 strand as a new structural determinant of neonicotinoid actions
Ihara, M., Okajima, T., Yamashita, A., Oda, T., Asano, T., Matsui, M., Sattelle, D.B., Matsuda, K.(2014) Mol Pharmacol 86: 736-746
- PubMed: 25267717 Search on PubMed
- DOI: https://doi.org/10.1124/mol.114.094698
- Primary Citation Related Structures:
3WTH, 3WTI, 3WTJ, 3WTK, 3WTL, 3WTM, 3WTN, 3WTO - PubMed Abstract:
Neonicotinoid insecticides target insect nicotinic acetylcholine receptors (nAChRs). Their widespread use and possible risks to pollinators make it extremely urgent to understand the mechanisms underlying their actions on insect nAChRs. We therefore elucidated X-ray crystal structures of the Lymnaea stagnalis acetylcholine binding protein (Ls-AChBP) and its Gln55Arg mutant, more closely resembling insect nAChRs, in complex with a nitromethylene imidacloprid analog (CH-IMI) and desnitro-imidacloprid metabolite (DN-IMI) as well as commercial neonicotinoids, imidacloprid, clothianidin, and thiacloprid. Unlike imidacloprid, clothianidin, and CH-IMI, thiacloprid did not stack with Tyr185 in the wild-type Ls-AChBP, but did in the Gln55Arg mutant, interacting electrostatically with Arg55. In contrast, DN-IMI lacking the NO2 group was directed away from Lys34 and Arg55 to form hydrogen bonds with Tyr89 in loop A and the main chain carbonyl of Trp143 in loop B. Unexpectedly, we found that several neonicotinoids interacted with Lys34 in loop G on the β1 strand in the crystal structure of the Gln55Arg mutant. Basic residues introduced into the α7 nAChR at positions equivalent to AChBP Lys34 and Arg55 enhanced agonist actions of neonicotinoids, while reducing the actions of acetylcholine, (-)-nicotine, and DN-IMI. Thus, not only the basic residues in loop D, but also those in loop G determine the actions of neonicotinoids. These novel findings provide new insights into the modes of action of neonicotinoids and emerging derivatives.
- Department of Applied Biological Chemistry, Faculty of Agriculture, Kinki University, Nara, Japan (M.I., Ta.O., T.A., M.M., K.M.); Institute of Scientific and Industrial Research, Osaka University, Ibaraki, Osaka, Japan (To.O.); Graduate School of Medicine, Dentistry and Pharmaceutical Sciences, Okayama University, Kita-ku, Okayama, Japan (A.Y.); and The Wolfson Institute for Biomedical Research, Department of Medicine, University College London, London, United Kingdom (D.B.S.) makoto_i@nara.kindai.ac.jp kmatsuda@nara.kindai.ac.jp.
Organizational Affiliation: