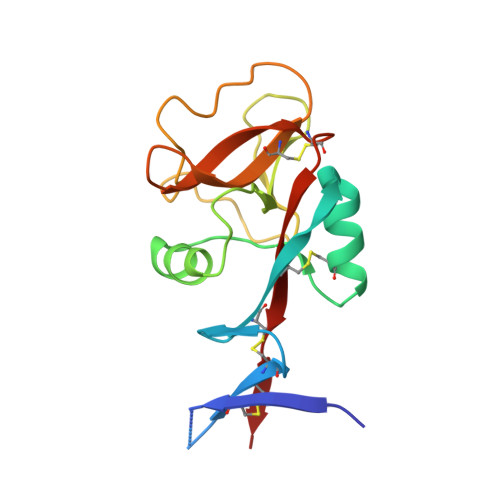
Structural analysis for glycolipid recognition by the C-type lectins Mincle and MCL
Furukawa, A., Kamishikiryo, J., Mori, D., Toyonaga, K., Okabe, Y., Toji, A., Kanda, R., Miyake, Y., Ose, T., Yamasaki, S., Maenaka, K.(2013) Proc Natl Acad Sci U S A 110: 17438-17443
- PubMed: 24101491 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1312649110
- Primary Citation Related Structures:
3WH2, 3WH3, 3WHD - PubMed Abstract:
Mincle [macrophage inducible Ca(2+)-dependent (C-type) lectin; CLEC4E] and MCL (macrophage C-type lectin; CLEC4D) are receptors for the cord factor TDM (trehalose-6,6'-dimycolate), a unique glycolipid of mycobacterial cell-surface components, and activate immune cells to confer adjuvant activity. Although it is known that receptor-TDM interactions require both sugar and lipid moieties of TDM, the mechanisms of glycolipid recognition by Mincle and MCL remain unclear. We here report the crystal structures of Mincle, MCL, and the Mincle-citric acid complex. The structures revealed that these receptors are capable of interacting with sugar in a Ca(2+)-dependent manner, as observed in other C-type lectins. However, Mincle and MCL uniquely possess shallow hydrophobic regions found adjacent to their putative sugar binding sites, which reasonably locate for recognition of fatty acid moieties of glycolipids. Functional studies using mutant receptors as well as glycolipid ligands support this deduced binding mode. These results give insight into the molecular mechanism of glycolipid recognition through C-type lectin receptors, which may provide clues to rational design for effective adjuvants.
- Laboratory of Biomolecular Science, Faculty of Pharmaceutical Sciences, Hokkaido University, Sapporo 060-0812, Japan.
Organizational Affiliation: