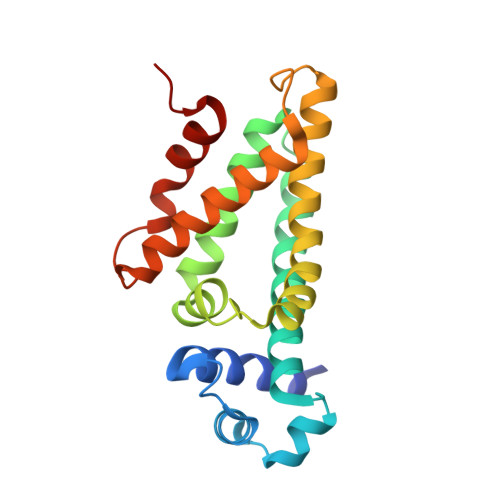
Structural characterization of a ligand-bound form of Bacillus subtilis FadR involved in the regulation of fatty acid degradation.
Fujihashi, M., Nakatani, T., Hirooka, K., Matsuoka, H., Fujita, Y., Miki, K.(2014) Proteins 82: 1301-1310
- PubMed: 24356978 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24496
- Primary Citation Related Structures:
3WHB, 3WHC - PubMed Abstract:
Bacillus subtilis FadR (FadR(Bs)), a member of the TetR family of bacterial transcriptional regulators, represses five fad operons including 15 genes, most of which are involved in β-oxidation of fatty acids. FadR(Bs) binds to the five FadR(Bs) boxes in the promoter regions and the binding is specifically inhibited by long-chain (C14-C20 ) acyl-CoAs, causing derepression of the fad operons. To elucidate the structural mechanism of this regulator, we have determined the crystal structures of FadR(Bs) proteins prepared with and without stearoyl(C18)-CoA. The crystal structure without adding any ligand molecules unexpectedly includes one small molecule, probably dodecyl(C12)-CoA derived from the Escherichia coli host, in its homodimeric structure. Also, we successfully obtained the structure of the ligand-bound form of the FadR(Bs) dimer by co-crystallization, in which two stearoyl-CoA molecules are accommodated, with the binding mode being essentially equivalent to that of dodecyl-CoA. Although the acyl-chain-binding cavity of FadR(Bs) is mainly hydrophobic, a hydrophilic patch encompasses the C1-C10 carbons of the acyl chain. This accounts for the previous report that the DNA binding of FadR(Bs) is specifically inhibited by the long-chain acyl-CoAs but not by the shorter ones. Structural comparison of the ligand-bound and unliganded subunits of FadR(Bs) revealed three regions around residues 21-31, 61-76, and 106-119 that were substantially changed in response to the ligand binding, and particularly with respect to the movements of Leu108 and Arg109. Site-directed mutagenesis of these residues revealed that Arg109, but not Leu108, is a key residue for maintenance of the DNA-binding affinity of FadR(Bs).
- Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto, 606-8502, Japan.
Organizational Affiliation: