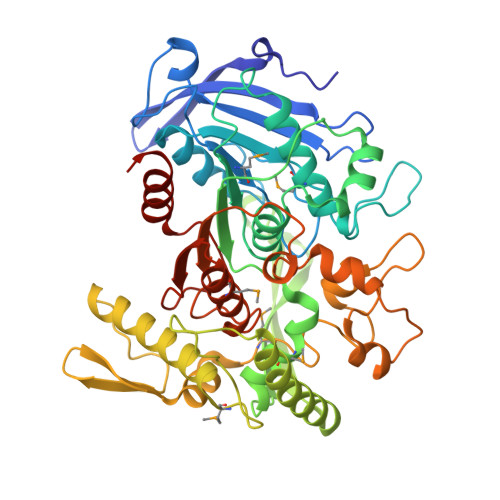
Crystallographic and mutational analyses of tannase from Lactobacillus plantarum.
Matoba, Y., Tanaka, N., Noda, M., Higashikawa, F., Kumagai, T., Sugiyama, M.(2013) Proteins 81: 2052-2058
- PubMed: 23836494 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24355
- Primary Citation Related Structures:
3WA6, 3WA7 - PubMed Abstract:
Tannin acylhydrolase (EC 3.1.1.20) referred commonly as tannase catalyzes the hydrolysis of the galloyl ester bond of tannins to release gallic acid. Although the enzyme is useful for various industries, the tertiary structure is not yet determined. In this study, we determined the crystal structure of tannase produced by Lactobacillus plantarum. The tannase structure belongs to a member of α/β-hydrolase superfamily with an additional "lid" domain. A glycerol molecule derived from cryoprotectant solution was accommodated into the tannase active site. The binding manner of glycerol to tannase seems to be similar to that of the galloyl moiety in the substrate.
- Department of Molecular Microbiology and Biotechnology, Graduate School of Biomedical & Health Sciences, Hiroshima University, Kasumi 1-2-3, Minami-ku, Hiroshima, 734-8551, Japan.
Organizational Affiliation: