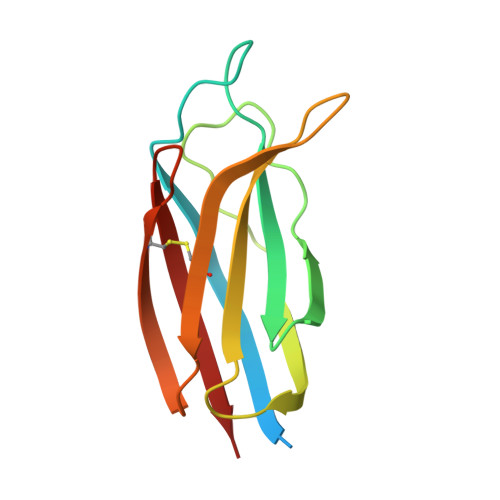
Defining the interaction of perforin with calcium and the phospholipid membrane.
Traore, D.A., Brennan, A.J., Law, R.H., Dogovski, C., Perugini, M.A., Lukoyanova, N., Leung, E.W., Norton, R.S., Lopez, J.A., Browne, K.A., Yagita, H., Lloyd, G.J., Ciccone, A., Verschoor, S., Trapani, J.A., Whisstock, J.C., Voskoboinik, I.(2013) Biochem J 456: 323-335
- PubMed: 24070258 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20130999
- Primary Citation Related Structures:
3W56, 3W57 - PubMed Abstract:
Following its secretion from cytotoxic lymphocytes into the immune synapse, perforin binds to target cell membranes through its Ca(2+)-dependent C2 domain. Membrane-bound perforin then forms pores that allow passage of pro-apoptopic granzymes into the target cell. In the present study, structural and biochemical studies reveal that Ca(2+) binding triggers a conformational change in the C2 domain that permits four key hydrophobic residues to interact with the plasma membrane. However, in contrast with previous suggestions, these movements and membrane binding do not trigger irreversible conformational changes in the pore-forming MACPF (membrane attack complex/perforin-like) domain, indicating that subsequent monomer-monomer interactions at the membrane surface are required for perforin pore formation.
- ††Crystallography, Institute of Structural and Molecular Biology, Birkbeck College, Malet Street, London WC1E 7HX, U.K.
Organizational Affiliation: